This is part of a series of posts about and leading up to the release of MacVector 12.5.
Assembler has always made it easy to assemble your sequencing projects. It hides the complicated algorithms and provides a point and click interface to show you the results. With the release of MacVector 12.5 the range of tasks that Assembler will perform has been greatly extended. Now with the addition of high throughput reference assembly using the popular Bowtie algorithm, Assembler can now support the alignment of many millions of NGS reads against genome sized references. Assembler will also generate output in the popular BAM format. Collections of contigs can be exported in FastQ format for additional analysis. Additionally the interface has been enhanced to increase the number of reads that can be submitted for de novo assembly. ..and to analyse your assemblies SNP detection and reporting has been enhanced with VCF output from Bowtie alignments and listing of all the codon and amino acid changes between the consensus and reference sequence alignments.
Here’s a selection of tasks that Assembler will make easy for you
De novo assembly of Sanger trace files
Add in your Sanger sequencing trace files, basecall the reads to improve accuracy then assemble using quality scores.
de novo short read Assembly
Add short reads from a variety of sources in FASTQ format as well as Sanger sequencing. Great for Hybrid assemblies
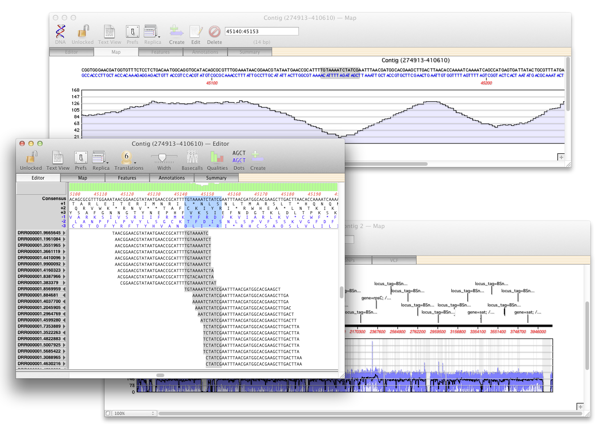
Reference assembly to identify SNPs in a bacterial/viral isolate
Add a Reference sequence and an NGS file(s) representing the sequence of an individual isolate and assemble using Bowtie. View a report listing all the potential SNPs based on the differences between the consensus and the reference. Be able to quickly identify the genes that the SNPs lie in and drill down to view the nucleotide and amino acid changes.
De novo bacterial assembly assisted by a Reference scaffold
Create a reference assembly using Bowtie, then take the individual contig consensus sequences along with all the input NGS Reads that did not assemble and assemble with a de novo assembler (Phrap) directly from within a new Assembly Project.
Assembly to multiple similar references
Take reference sequences from a series of closely related strains of virus or bacteria. The reads come from a single isolate. Use Bowtie to assemble these against the collection of References to determine which is most closely related (or identical) to the isolate.
Assembly to multiple dissimilar references to identify SNPs
Essentially similar to the bacterial SNP assembly, but using yeast or some other organism with multiple genomes (or even some bacteria that have multiple chromosomes or large plasmids).
Exome (transcriptome) sequencing)
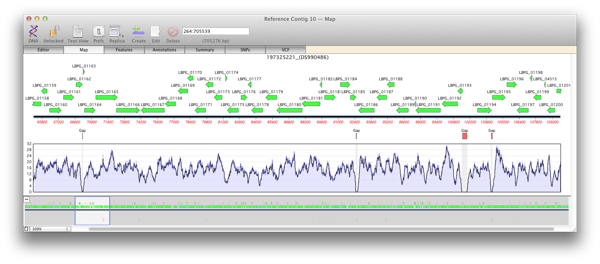
Use the genomic sequence of an organism as a reference and align reads from sequenced mRNA, cDNA or total RNA of that same organism. Uses are splice site junction identification, novel gene identification amongst many others.
..and remember if you purchase an upgrade or a new license before the release of MacVector 12.5 you will get a 10% discount and a free upgrade to MacVector 12.5 when it is released. Contact Sales@macvector.com and quote “MV12510” for the discount.
Technorati Tags: bowtie, MacVector
