Editing and Analysis
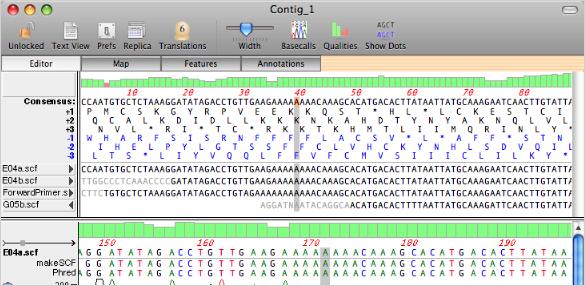
You can edit the sequences in a contig and the consensus will be updated automatically. MacVector follows the phred/phrap quality value rule where edited residues are given a quality value of 99 to indicate they have been assigned by a user. These are shown in blue in the quality display. You can view a variety of statistics on the composition of the contig in the annotations window.
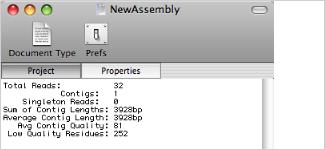

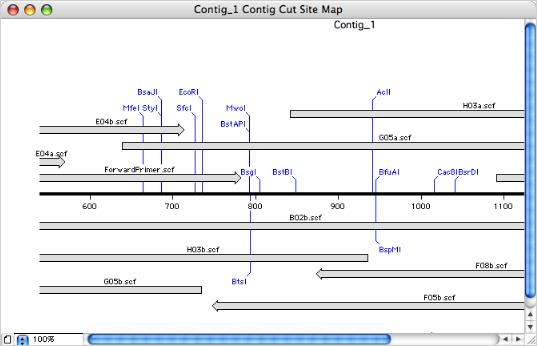
You can invoke any MacVector DNA analysis algorithm from the contig editor window, including online NCBI blast searches. Any gaps in the consensus are removed before the analysis is performed. This lets you scan the contig for restriction enzyme sites, then edit the consensus and rescan without having to export the consensus sequence or switch to a different module. The consensus sequence, without gaps, can be saved to disk as a single sequence at any time. The sequence can be saved in any format supported by MacVector and retains a list of the individual reads used to generate the consensus.

|
2x.png)

2x.png)
