Category: Techniques
-
Use a right-click in the Editor tab to see if your contig can be circularized
MacVector incorporates no less than THREE different de novo assemblers, phrap, velvet and SPAdes. While all are great assemblers, with each having their own specific advantages, none of them will generate a circular sequence from input reads. However, MacVector also includes a tool to help you with this. If you are assembling reads representing plasmid…
-
Import Multi-Sequence Genbank Files into an Assembly Project for easy access to Features
There are many genomes in the Genbank database that cannot be downloaded as single annotated sequences. These might be large multi-chromosome eukaryotic genomes, but, increasingly, partially sequenced bacterial chromosomes where the major contigs have been annotated using the NCBI annotation pipeline. Typically, when you encounter these, there are options to download annotated versions of these…
-
Optimizing Align To Folder Parameters for use with NGS Data
You can use the Database | Align To Folder function to scan large fasta or fastq files containing NGS data to find and retrieve just those reads that match a specific target sequence. The search is aware of paired-end reads, so when you retrieve hits, both reads of a pair will be saved into a…
-
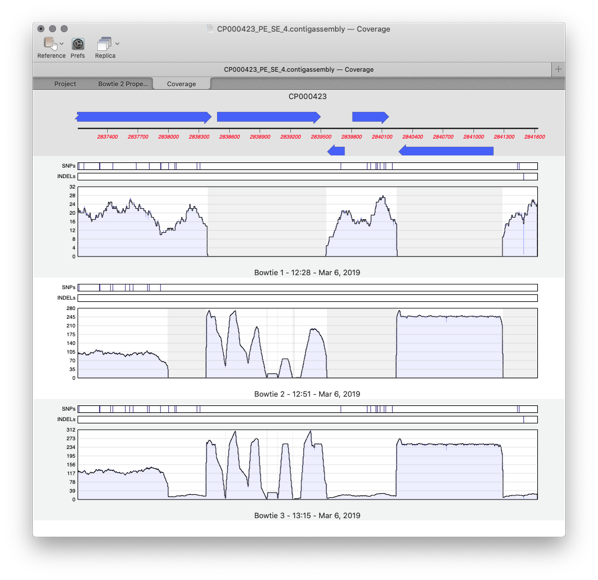
Comparing multiple reference assemblies
MacVector 17 has a greatly improved Assembly Projects manager, for better organization of multiple sequencing datasets, multiple references sequences and repeated jobs. Every time you run a new assembly job (either a reference assembly or de novo). A new job object is created in the Assembly Project window contains resulting contigs and any unaligned reads…
-
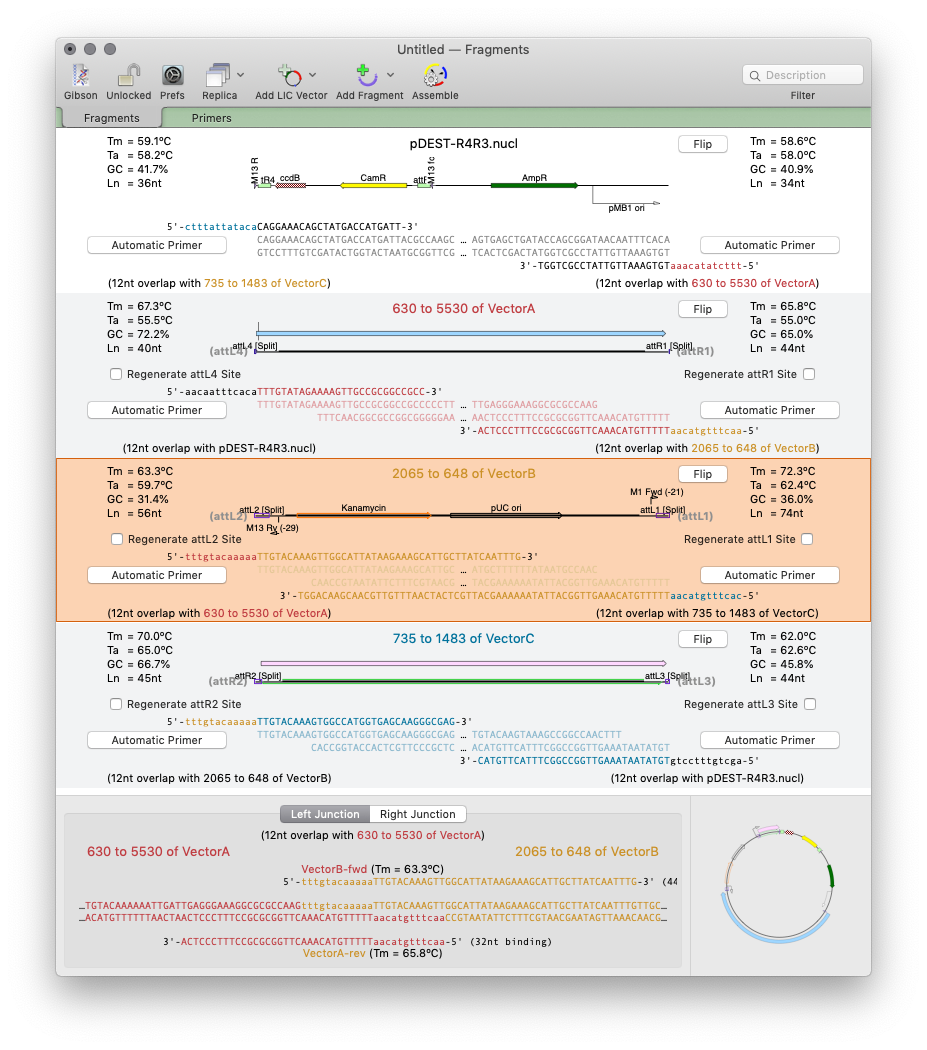
Designing primers for Gibson Assembly with MacVector 17
MacVector 17 has a completely new tool for automated design of ligase-independent cloning strategies. The tool supports 5’ exonuclease driven Gibson assembly as well as the T4 DNA Polymerase 3’ exonuclease “Ligase Independent Cloning” approach. MacVector can automatically design primers when you specify fragments and vectors to use. You can provide custom primers (manually or…
-
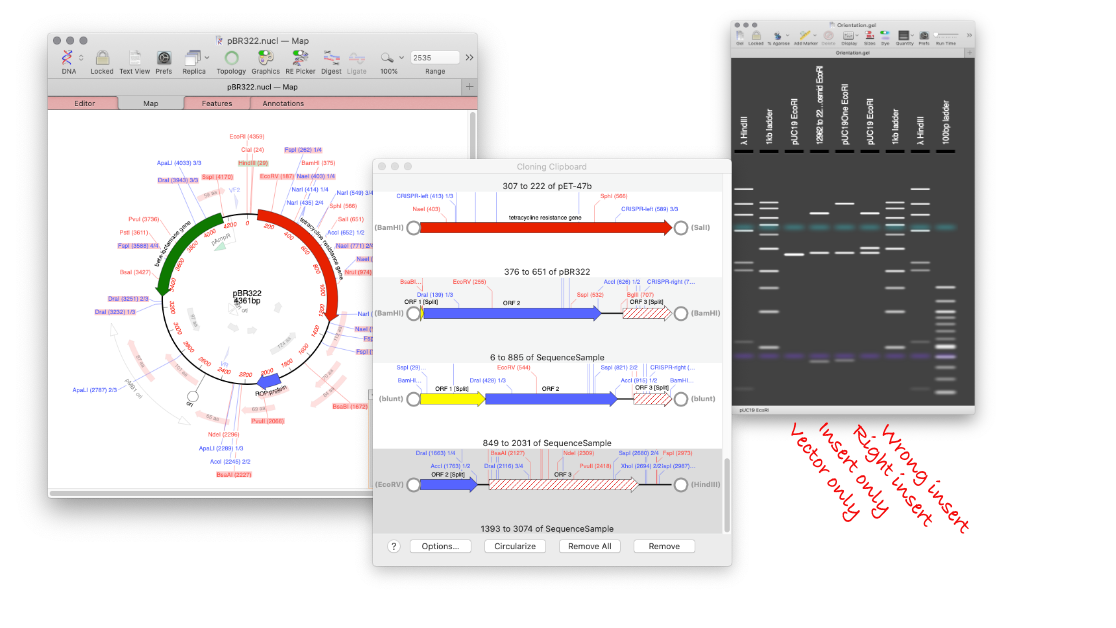
Using MacVector’s Agarose Gel tool to design a digest to screen minipreps after a ligation.
How to design a digest to screen minipreps after a ligation.
-
RNASeq Expression Analysis with MacVector and Assembler
If you have the Assembler module, MacVector can align millions of NGS reads from RNASeq experiments against large genomes and generate a coverage table displaying the relative expression levels of every gene in a genome. The key to this functionality is that you must have a reference genome with genes annotated as CDS or gene…
-
How to split large fastq files for more manageable assemblies
We’ve previously discussed how important it can be to make sure you are using the appropriate number of fastq reads from an NGS experiment to ensure you obtain the results you are looking for. Using too many reads can confuse algorithms with the massive coverage increasing mis-assemblies due to background errors in the reads. In…
-
Workflows on designing, testing and storing primers in MacVector
MacVector is the best PCR Primer design application on macOS. It has many innovative primer tools to make designing, analyzing and cataloging your primers or oligos easy. Here are a few typical workflows. Designing primers Amplifying a gene You can design a set of primers to amplify a gene in as little as three mouse…
-
Assembling sequencing data with MacVector and Assembler
Update: As of September 2024 Assembler will be added at no cost when you upgrade or renew maintenance. Contact Support if you have not yet been upgraded. To assemble various types of sequencing reads, follow these steps. Then follow one of the following: To create a de novo assembly from Sanger reads To create a…
