Category: Tips
-
MacVector video tutorials on You Tube
There are short video tips on using MacVector on our blog and our YouTube channel. Each one is less than two minutes and generally shorter than that! Please subscribe so you don’t miss any! The latest screencasts include quickly annotating a gene to a sequence, confirming a small sequencing project against a reference and checking…
-
Use the Replica Button For Synchronized Views
Most primary MacVector windows (Nucleic Acid Sequence, Protein Sequence, Multiple Sequence Alignment, Align To Reference, Contig Assembly etc.) have a Replica toolbar button. If you click that button, a second window will open, potentially set to a different tab. The key to this functionality is that the two windows are linked – any selections you…
-
How to Identify Bacterial Promoters Using MacVector
MacVector’s Subsequence tool is a very flexible search function that can be used for a variety of tasks. MacVector itself has a built-in variant of the function for maintaining and search primer databases (Analyze | Primer Database Search…). Each entry in the file MacVector uses as a source of subsequence data can have up to…
-
Import Multi-Sequence Genbank Files into an Assembly Project for easy access to Features
There are many genomes in the Genbank database that cannot be downloaded as single annotated sequences. These might be large multi-chromosome eukaryotic genomes, but, increasingly, partially sequenced bacterial chromosomes where the major contigs have been annotated using the NCBI annotation pipeline. Typically, when you encounter these, there are options to download annotated versions of these…
-
Opening multiple sequences as alignments or individual sequences
Many sequence formats contain multiple concatenated sequence entries. For example FASTA and Genbank are two formats capable of storing multiple individual sequences. By default MacVector will treat such sequences as alignments and open them in the Multiple Sequence Alignment editor. Most users who want to open such a file do want to see an alignment.…
-
Restoring file associations when MacVector no longer opens your sequences
Macs are pretty good at choosing the right application to open a document. For example when you double click on a .nucl document then it will open in MacVector. However, sometimes this file association breaks. Applications should coexist peacefully on a Mac, but sometimes a misbehaving app will corrupt these file associations and you will…
-
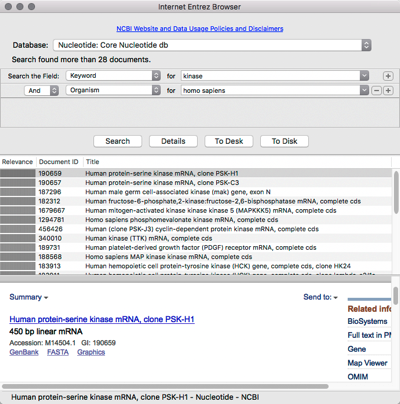
Searching and downloading sequences from Entrez
MacVector has integrated connectivity to the NCBI BLAST and Entrez databases. You can directly search Entrez for DNA or Protein sequences based on features, authors, keywords etc and directly download them into MacVector, complete with all features and annotations.
-
Use a right-click in the Contig Editor tab to see if your contig can be circularized
MacVector 16 incorporates no less than THREE different de novo assemblers, phrap, velvet and SPAdes. While all are great assemblers, with each having their own specific advantages, none of them will generate a circular sequence from input reads. However, MacVector 16 also includes a new feature to help you with this. If you are assembling…
-
Downloading hits from the MacVector BLAST Map results tab
MacVector’s BLAST Map results tab (added in MacVector 15.5) is a unique interface for examining the annotations around hits to a query sequence. Each pane in the display represents a High Scoring Segment Pair, as seen in the BLAST Aligned Sequence tab. At the lower left corner of each pane is a Download button –…
-
Turning on/off the SCAN FOR missing Features and ORFs
If you’re running MacVector 15.5 or later then you will have noticed extra features annotated to your sequences. These are from the Scan for ORFs tool (added in MacVector 15.5) and the Scan For Missing Features (added in MacVector 16) tools that automatically scan every DNA sequence window for open reading frames, missing annotations and…
