Align to Reference - cDNA Alignment
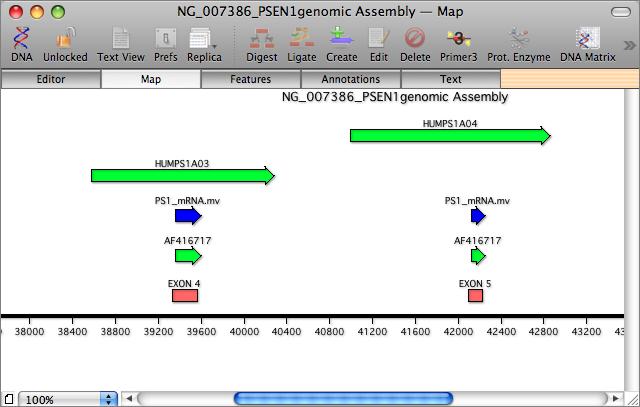
MacVector has a unique Align to Reference interface that lets you align one or more files against a reference sequence. There are two main uses for this: Sequence Confirmation is similar to sequence assembly, except that it requires the use of a known reference sequence as a scaffold. The second use is cDNA Alignments, which allows you to align mRNA, cDNA or EST sequences against a genomic template.
There is a detailed Sequence Confirmation Tutorial that provides far more information on this functionality that can be downloaded here. The trial version of MacVector includes all of the sample files you will need to follow the tutorial.
Align to Reference Editor
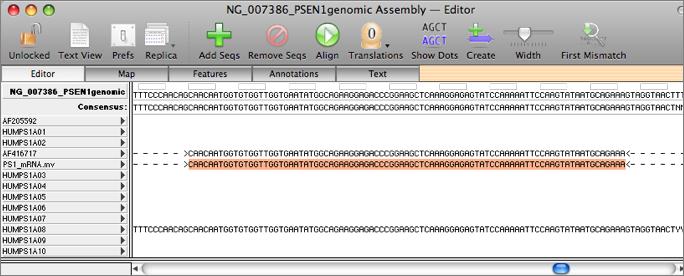
The key to this functionality lies in the main Align to Reference editor. You start with a reference sequence. This is displayed along the top of the window. You can then add one or more sequences to the window by clicking on the "+" button. The imported sequences are displayed below the reference sequence. Initially, the sequence residues are displayed in italics to indicate they have not been aligned.




|

2x.png)


2x.png)
