You can use the Analyze | Align to Reference function to align other sequences (Sanger chromatograms, plain sequences or even NGS data collections) against a reference. Once aligned, the Editor lets you perform all the usual editing functions using an “overwrite” mode – select the residue you want and type the new residue to replace it. Typing a “space” or a gap “-” will delete the residue. However, there are a number of other editing functions you should be aware of:
- Hold down the [option] key and type a residue to insert a residue, rather than overwrite. The new residue inserts before the selected base.
- Hold down [option] and type [delete] to delete a residue rather than replace it with a gap. This will cause the residues to the right of the deletion to slide to the left by one character.
- To “nudge” an entire read, simply select the “name” of the read in the top left pane and press the [left] or [right] arrow keys.
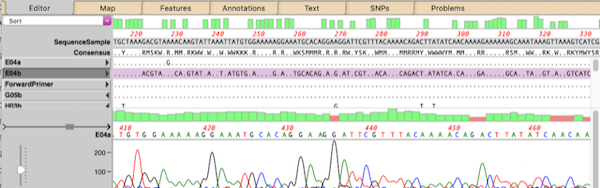
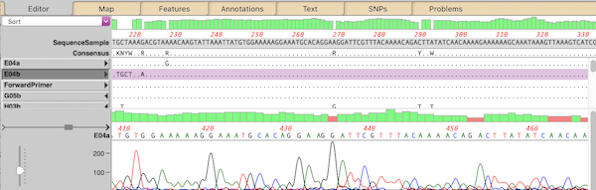
- There are a large number of editing functions that can be accessed through a context-sensitive right-click menu. MacVector 18 introduced a new Extra “hamburger” menu that shows the same list:

- Most of these operate on the entire set of selected read(s) and the actions are fairly self-explanatory. However, some may need additional explanation:
- (i) Select Matching Pairs – primarily relevant for NGS data, this will attempt to select the corresponding mate for paired-end reads, based entirely on the name.
- (ii) Select Overlapping Reads Containing Selected Sequence – if you select a short sequence in one of the reads, perhaps containing a SNP, this will select all of the sequences that contain that same SNP.
- (iii) Close Gaps by Deleting Residues – with large alignments, you may be distracted by a column of mostly gaps due to e.g. one read out of 100 having an additional inserted residue. This function removes these occasional insertions to clean up the alignment.
- (iv) Extend Reference with Selected Read – only works if a single read is selected that extends past either end of the reference sequence. Useful for building up “patch” sequences when trying to resolve repeats during genome assemblies.

