The Quicktest Primer interface is highly interactive and the display shows restriction enzymes and the amino acids sequences of CDS features in the region of interest. Here’s a ~25nt primer aligned against a parental sequence. Restriction enzyme recognition sites are shown in black text. “One-out” sites (e.g. a 5 out of 6 match) are shown with an asterisk. Those colored green would NOT change the CDS amino acid sequence if the mismatched residue was changed to create a complete recognition sequence. Conversely, red colored one-out sites WOULD change the amino acid sequence.
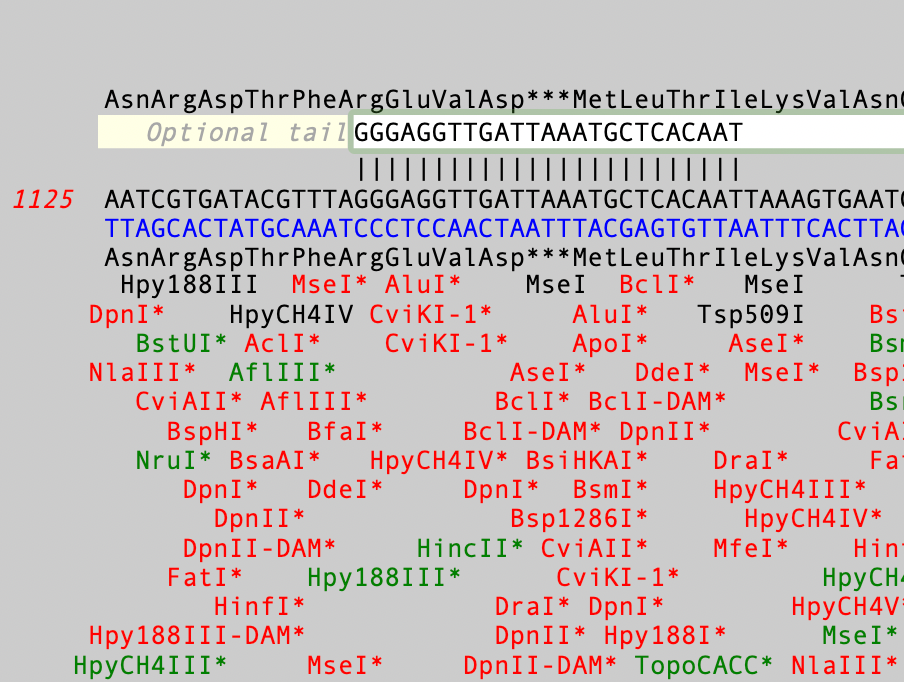
You can interactively view the cut site for any enzyme by moving the mouse pointer over a restriction site;
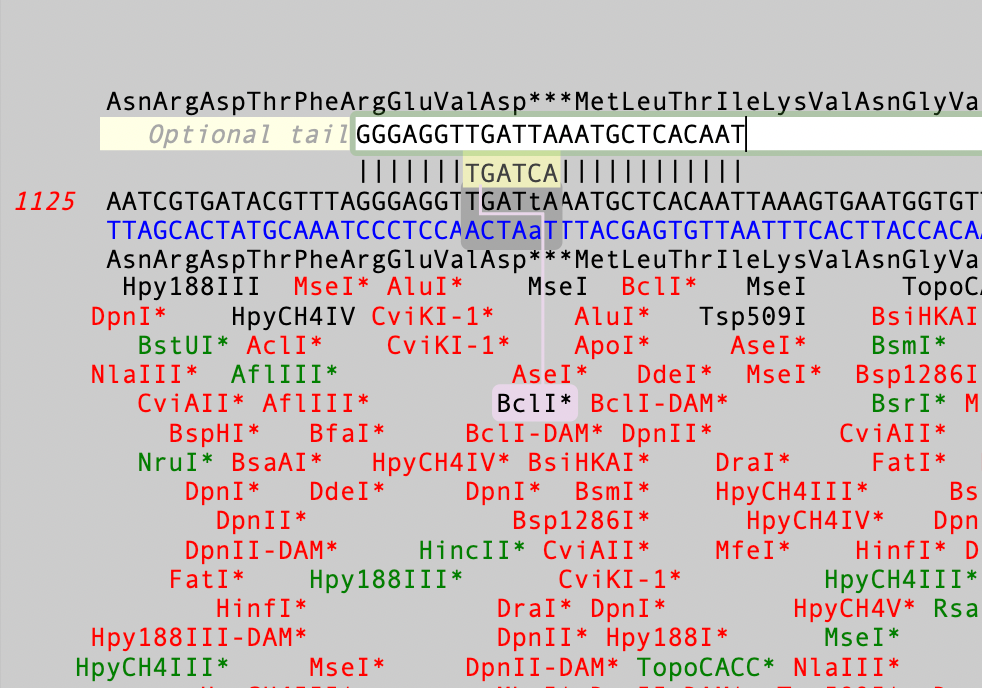
In this case you can see the TGATCA BclI site and, if you look closely, the one-out mismatched residue is shown in lower case i.e. TGATtA.Then, if you mouse-click and hold on a site, the display changes to show you what effect changing that mismatched residue would have on the primer and on the amino acid sequence.
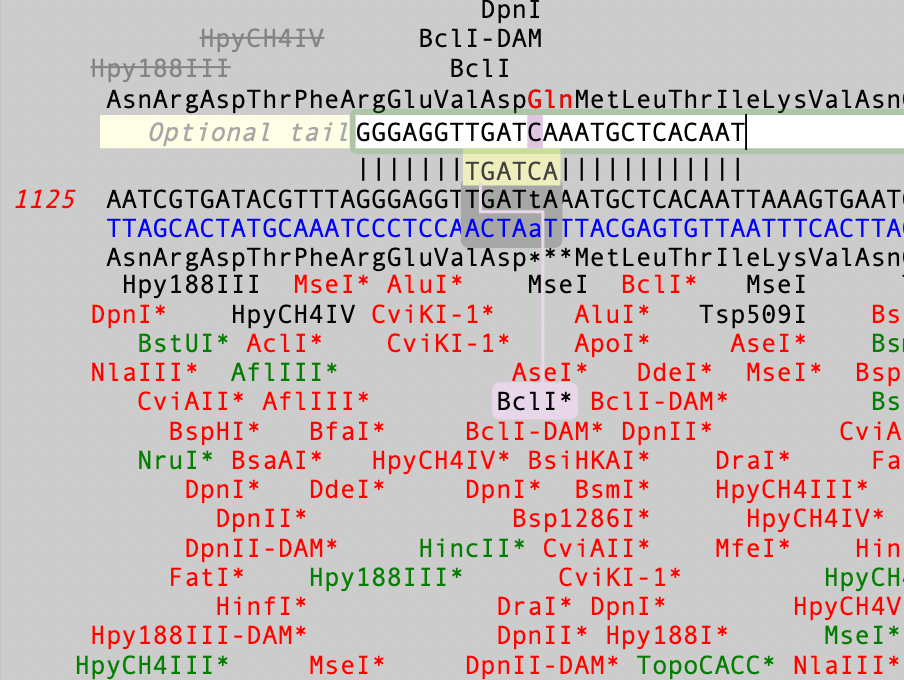
In this case, the stop codon is changed to a glutamine codon and a BclI site is created. Finally, if you double-click on a site, a permanent change is made to the primer sequence.For more information on the use of the Quicktest Primer interface, take a look at the Primer Design tutorial.
