Author: Chris
-
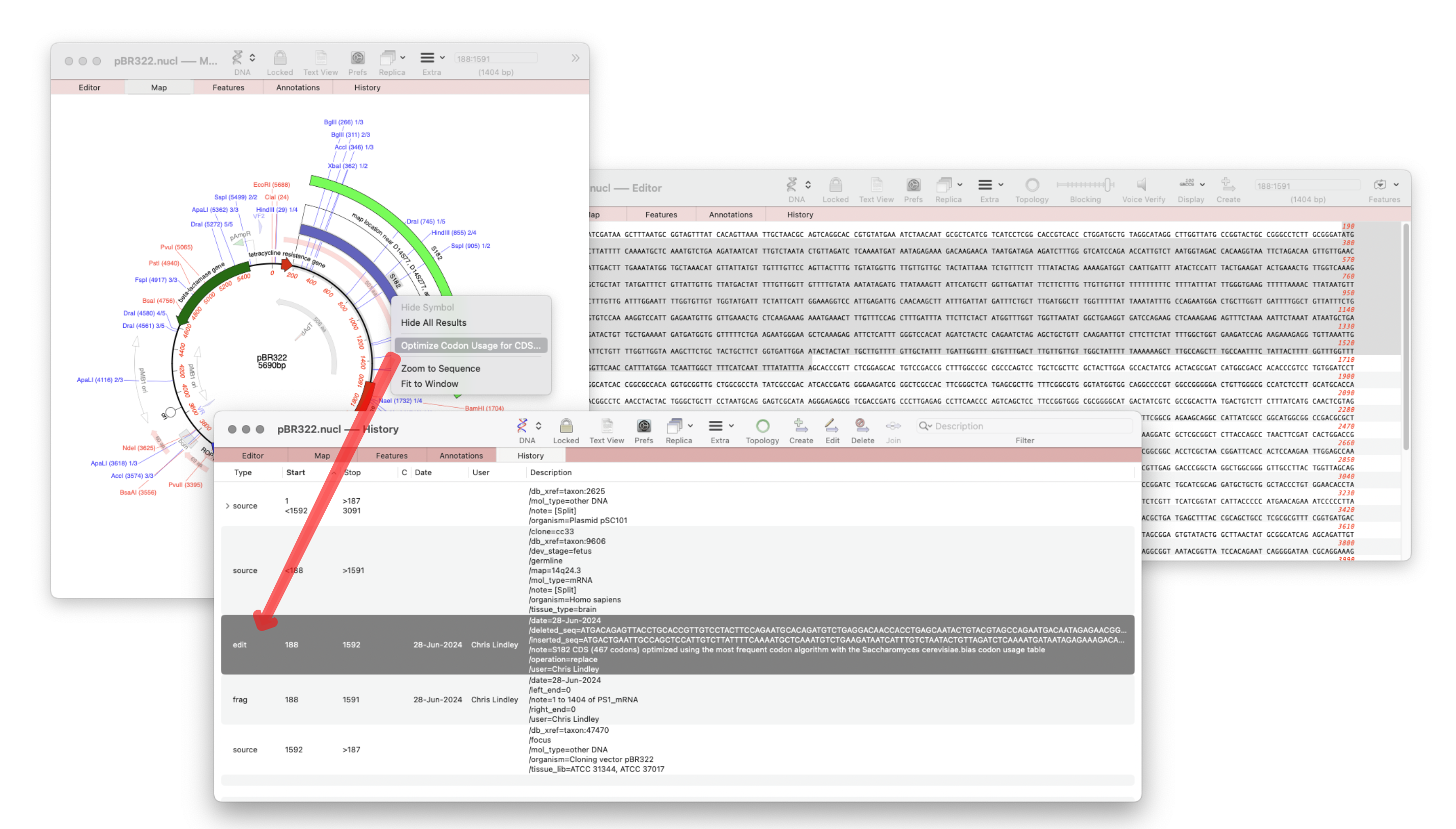
MacVector 18.7 – View the history of your plasmid
The recently released MacVector 18.7 has a new History tab in the Single sequence editor that shows the editing history of your DNA sequences Since the introduction of MacVector’s Cloning Clipboard, all cloning actions (such as ligating a digested fragment into a vector) create a /FRAG feature that records the source of the ligated fragment, the restriction enzymes used to digest it (and…
-
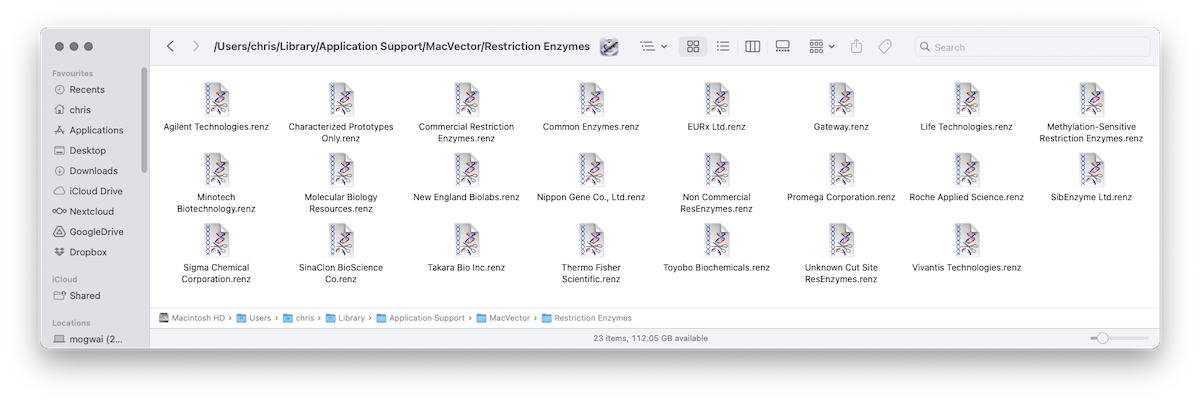
MacVector 18.7 – Change in Default Restriction Enzyme File Location
One change in MacVector 18.7, that will improve installation on multi user Macs, is that by default MacVector now stores restriction enzyme files in the user’s home folder. Since it’s the user’s home folder, it will always be writeable, even if the user does not have Administrator access to the machine. Additionally for a Mac…
-
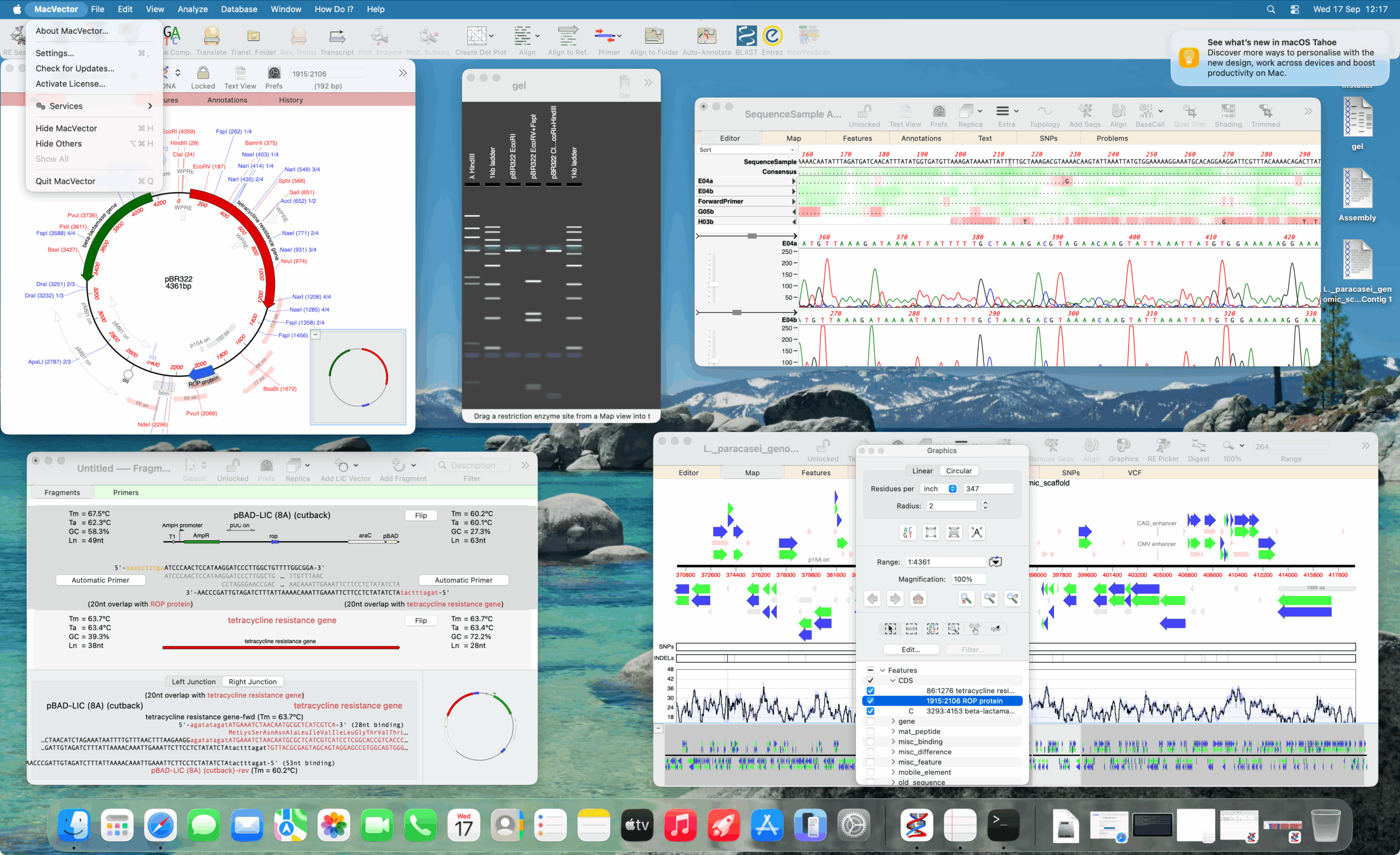
MacVector 18.7.2
MacVector 18.7.2 update is now out We’ve just released another minor update to MacVector 18.7. Sorry to follow an update with another update so quickly but we discovered two minor, but annoying bugs. You will be prompted to update over the next few days (unless you have turned off update notifications). However, you can also…
-

MacVector 18.7.1 update is now out
We’ve just released a minor update to MacVector 18.7. You will be prompted to update over the next few days (unless you have turned off update notifications). However, you can also run MACVECTOR | CHECK FOR UPDATES.. or download the new version directly Changes for MacVector 18.7.1 macOS Support This release now supports all macOS…
-
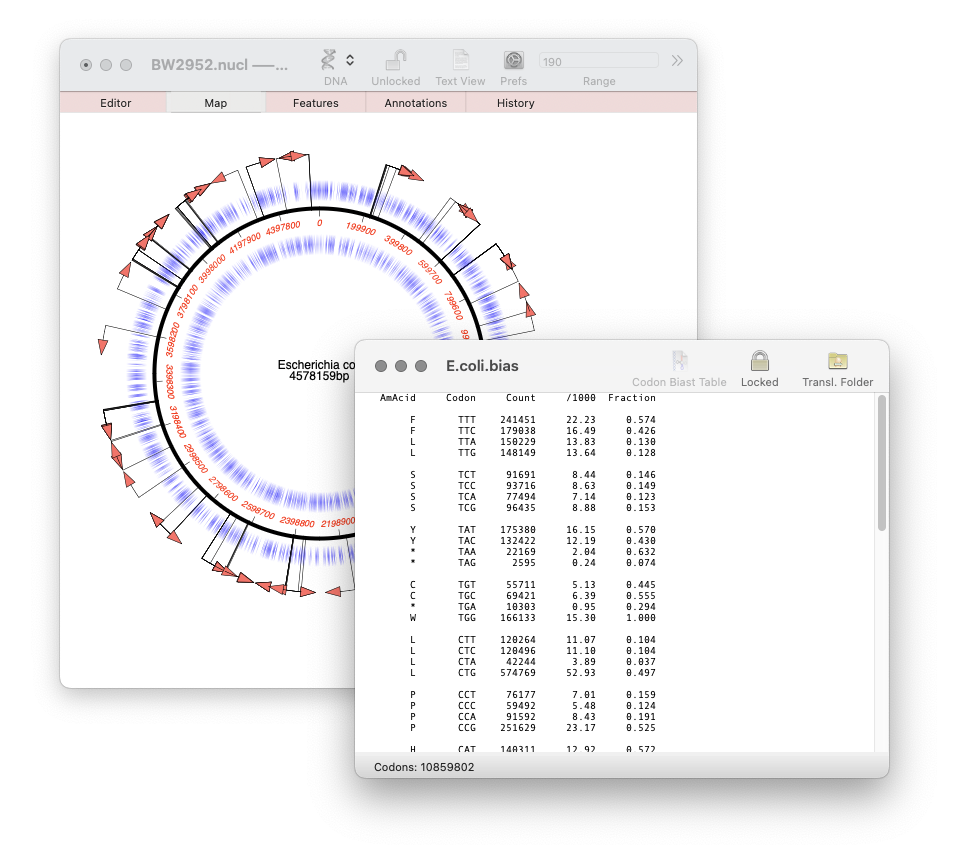
MacVector 18.7: Generating custom Codon Usage Tables (CUT) from your own sequences.
Our latest release, MacVector 18.7, has a new Codon Usage Table viewer. You can use this to generate your own codon usage table (CUT or .bias) files. You can use codon usage tables to optimize codon usage of CDS features for enhanced expression in a different organism. They can also be used in the Nucleic Acid Toolbox to predict protein coding…
-
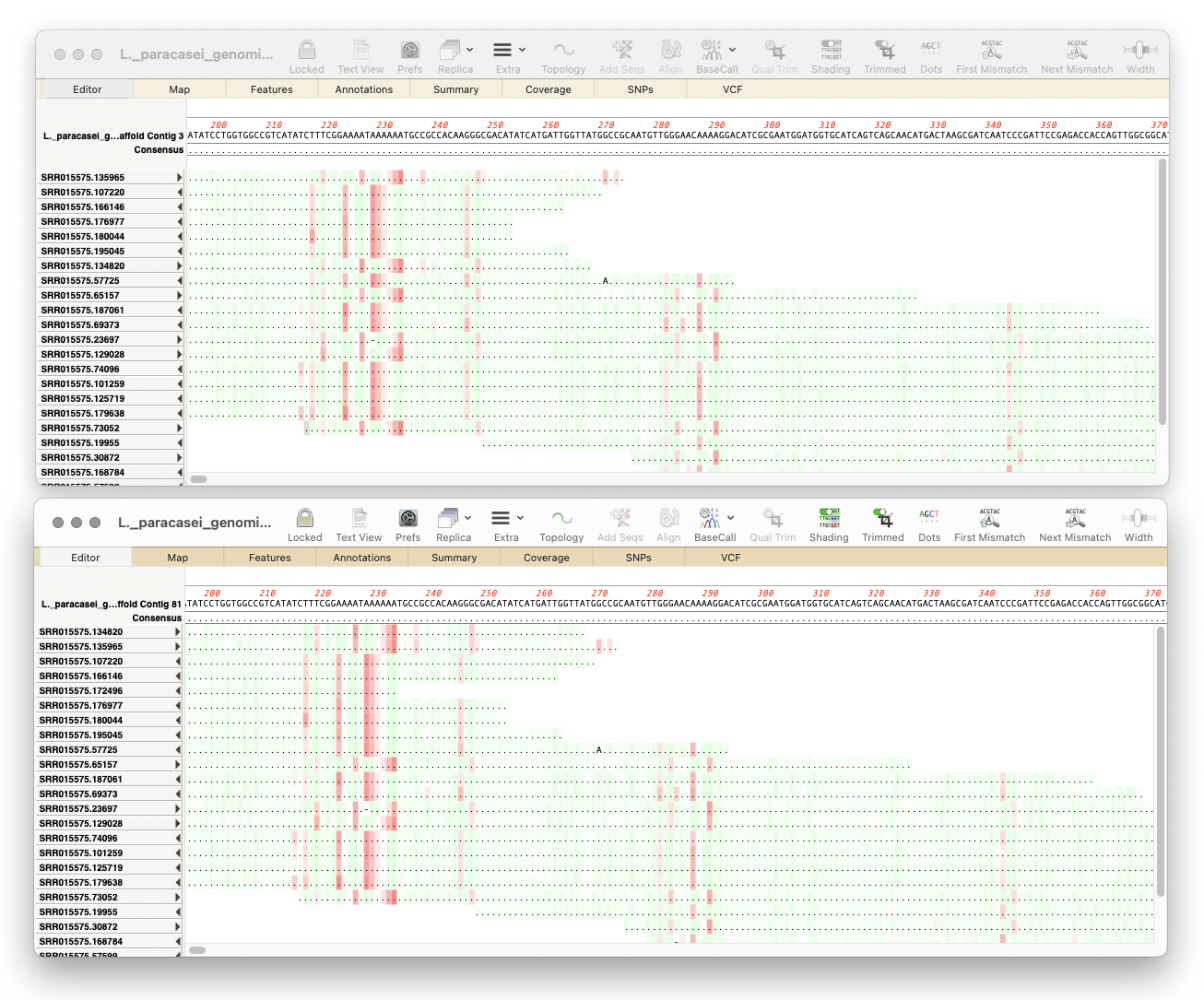
MacVector 18.7: Long-Read Reference Alignments using minimap2
Our latest release, MacVector 18.7, sees the addition of Minimap2 to Assembler’s sequencing toolkit. So if you have the Assembler module, you can now map noisy long-read data from Pacific Biosciences or Oxford Nanopore to one or more genomes. Minimap2 is a reference assembler similar to Bowtie2. But whereas Bowtie2 excels at mapping “short reads” (500nt or less) to…
-

MacVector 18.7 has just been released.
MacVector 18.7 has just been released. If you are eligible for this release you will be prompted to upgrade, otherwise go to MACVECTOR | CHECK FOR UPDATES… and follow the prompts to be automatically upgraded. If your license is not eligible then why not upgrade? Overview MacVector 18.7 introduces a History tab to track the…
-
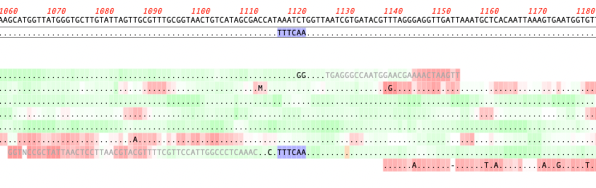
MacVectorTip: Quality scoring of manual edits to your contigs.
Quality scoring of Assemblies and Align to Reference alignments can be visualized directly on the sequence. Residues can be shaded according to their quality scores. These can be displayed anywhere quality values are available, including de novo and reference assemblies in Assembler and Align to Reference alignments. The intensity of the shading of residues indicates…
-
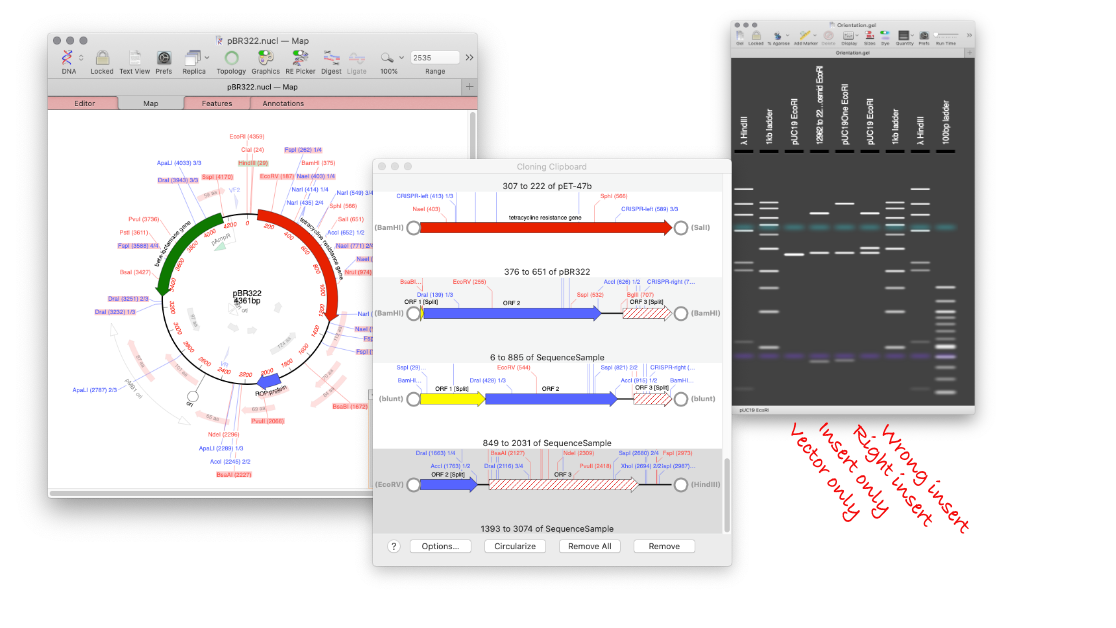
How to design a digest to screen minipreps after a ligation.
MacVector’s Agarose Gel tool can be used to quickly design a restriction digest to screen minipreps following a ligation. Replicate your ligation in MacVector. Create your agarose gel with the correct insert and a vector only lane. Undo the ligation, and repeat with the wrong orientation. Now you will end up with an Agarose Gel…
-
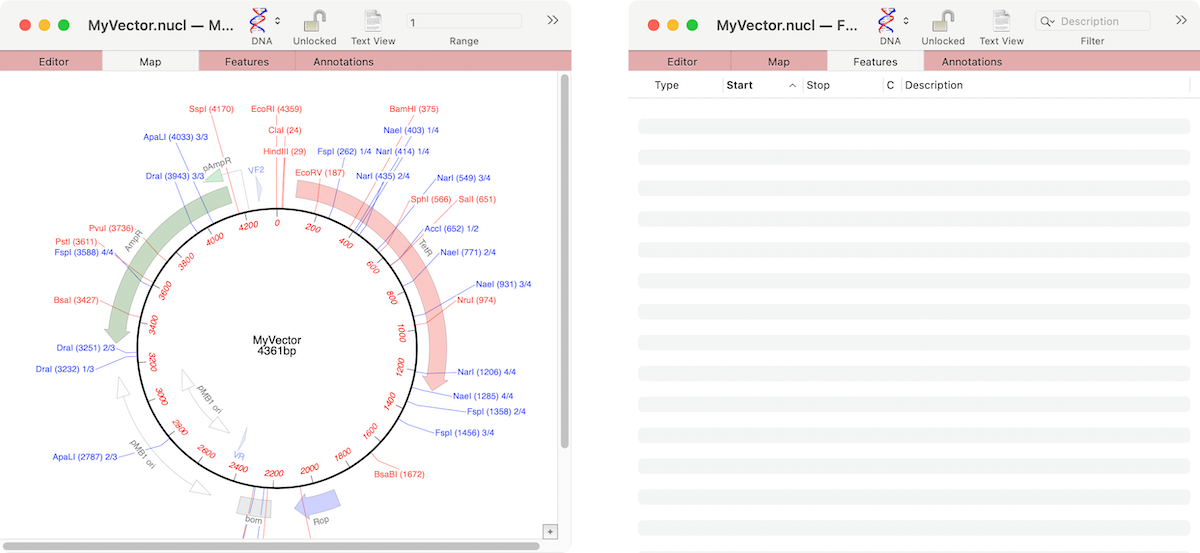
MacVectorTip: Grayed out graphics indicate Missing Features
If the graphics in a nucleic acid sequence Map tab appear somewhat “washed out” it is because the graphic items represent common features that MacVector has found that are not annotated on the sequence. For example, here are the Map and Feature tabs of an unannotated cloning vector; You can see a number of features…
