Category: Tips
-
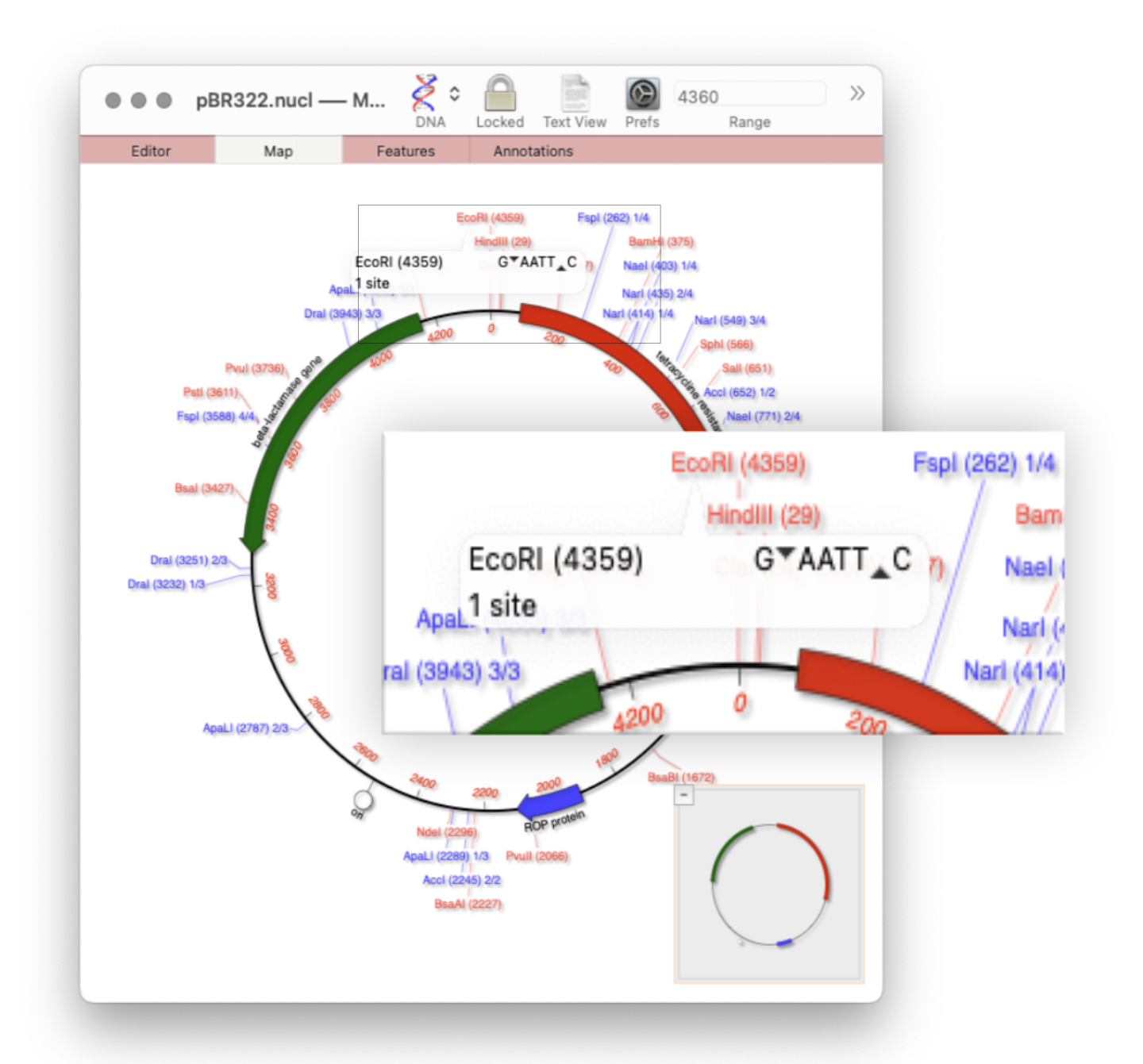
MacVectorTip: Restriction enzyme sites and tooltips
Quickly viewing the recognition sequence and cut site of a restriction site is very easy in the Map tab. By default MacVector’s Scan DNA For… tool will automatically display restriction enzyme recognition sites in the Map tab. If you hover your mouse over a restriction site, a tooltip will show you the restriction enzyme recognition site, the location of the cut…
-
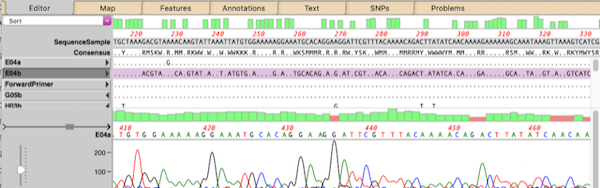
MacVectorTip: Advanced Align to Reference Editing
You can use the Analyze | Align to Reference function to align other sequences (Sanger chromatograms, plain sequences or even NGS data collections) against a reference. Once aligned, the Editor lets you perform all the usual editing functions using an “overwrite” mode – select the residue you want and type the new residue to replace…
-

MacVectorTip: How to Toggle Between Single-Letter and Three-Letter Amino Acid Translation Code
Many views in MacVector display amino acid translations above or below DNA sequences. Typically, these are from CDS features, but can also be the 3/6 frame translation of a sequence. You can display the amino acids as either the single-letter code, or as the three-letter code. You can toggle the setting using the MacVector | Preferences ->…
-
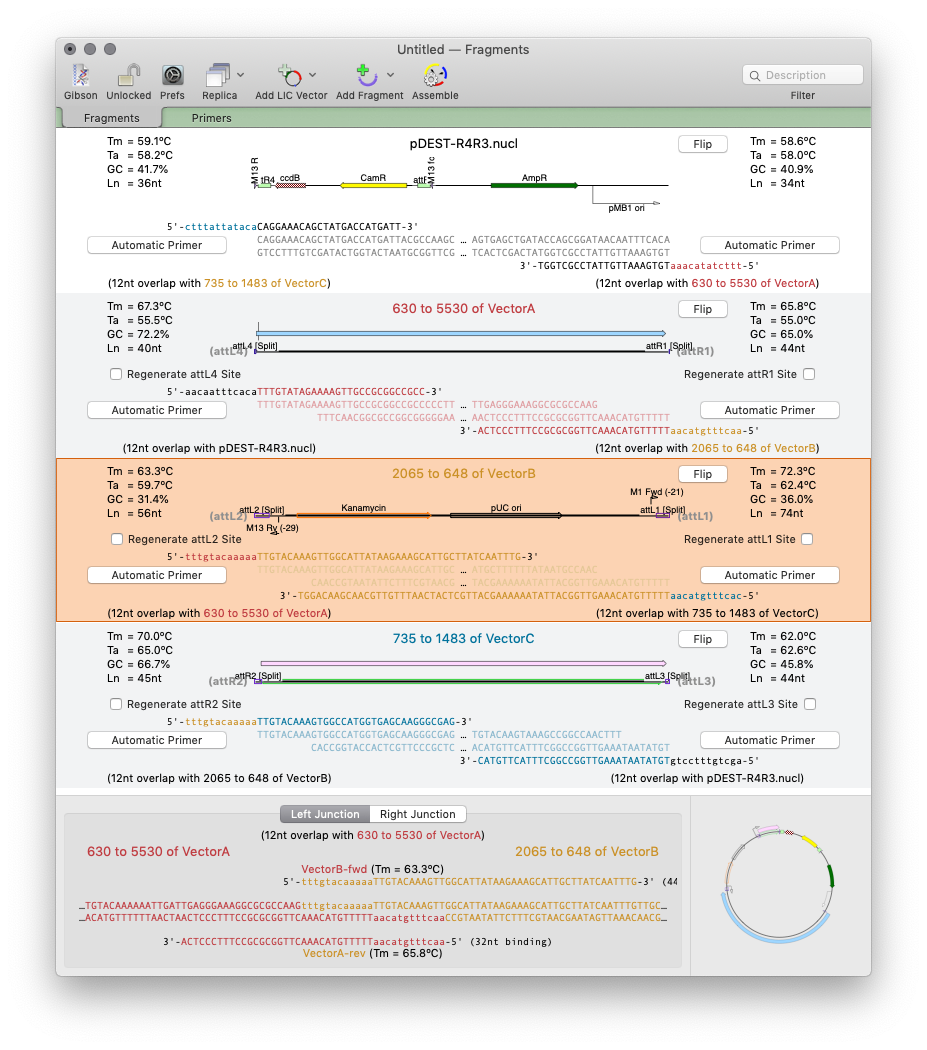
MacVectorTip: Drag and drop in the Gibson Assembly window
Over the years, we have added a lot of drag and drop functionality to MacVector. Of course, as with any application, it is not always obvious that you can drag and drop to accomplish tasks because you literally have to drag and drop to discover you can do it at all! So here is the first of…
-
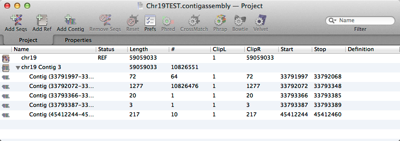
MacVectorTip: organizing assemblies and sequencing datasets with the Assembly Project manager.
MacVector’s Assembly Project manager helps you organize multiple sequencing datasets, multiple reference sequences and repeated assemblies. You can store multiple assembly jobs in a single Assembly Project and directly compare multiple runs of the same dataset to determine the best assembly parameters. You can also compare different sequencing datasets assembled against the same reference sequence…
-
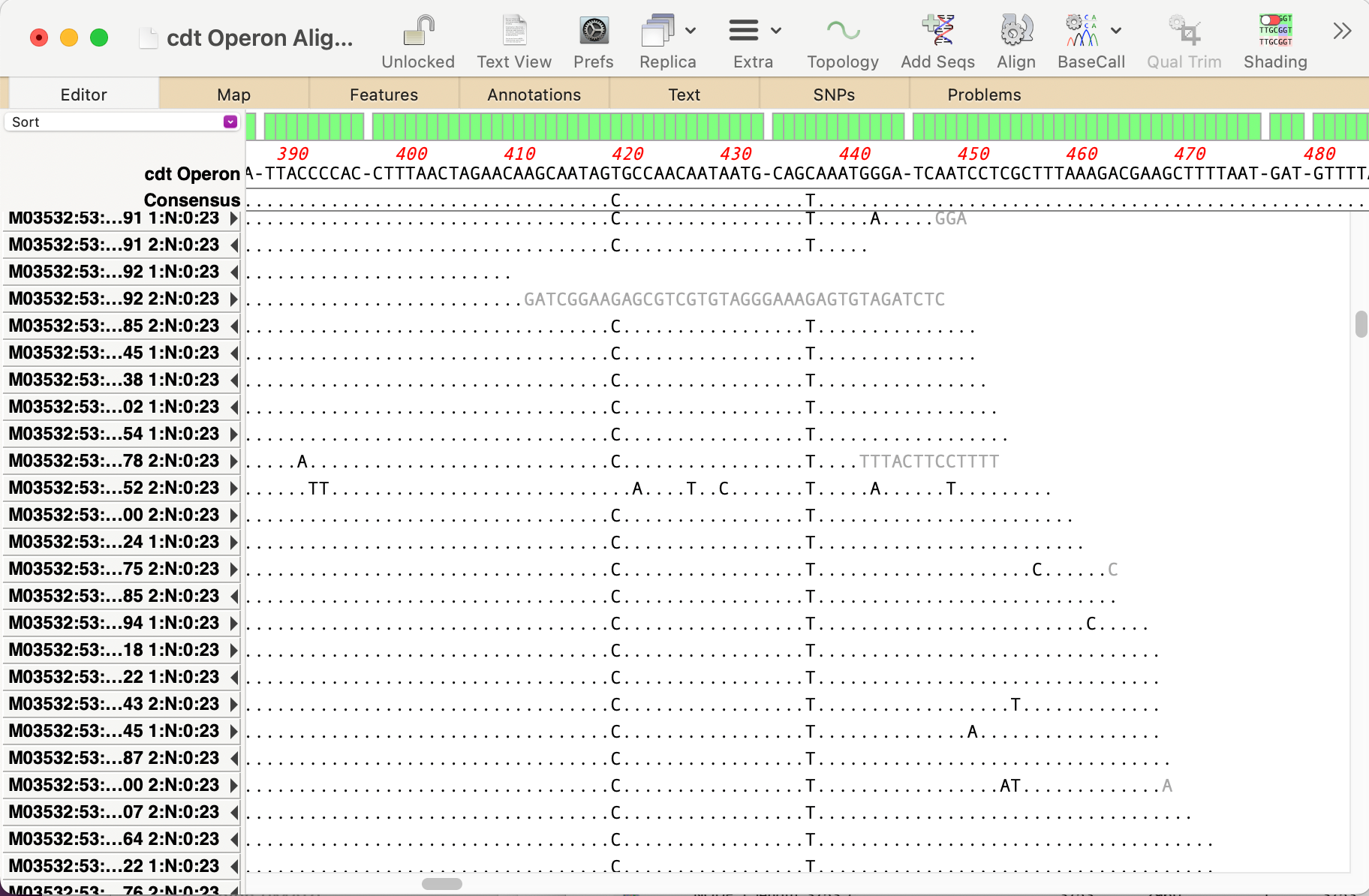
MacVectorTip: Identifying, Selecting and Assembling NGS reads with a variant genotype
When analyzing/assembling/aligning NGS data, there are many scenarios where you might want to separate out the reads representing different genotypes or variant sequences. MacVector makes this very easy. Take a reference sequence and choose Analyze->Align to Reference. Now click the Add Seqs button and select and add your NGS data files. NOTE: if your reference…
-
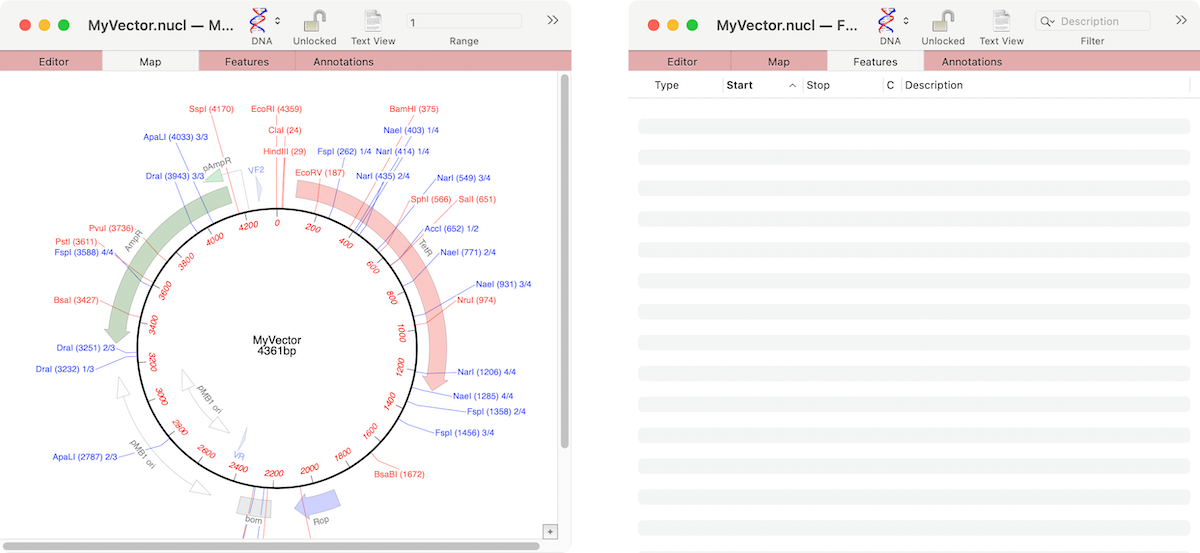
MacVectorTip: Grayed out graphics indicate Missing Features
If the graphics in a nucleic acid sequence Map tab appear somewhat “washed out” it is because the graphic items represent common features that MacVector has found that are not annotated on the sequence. For example, here are the Map and Feature tabs of an unannotated cloning vector. You can see a number of features…
-
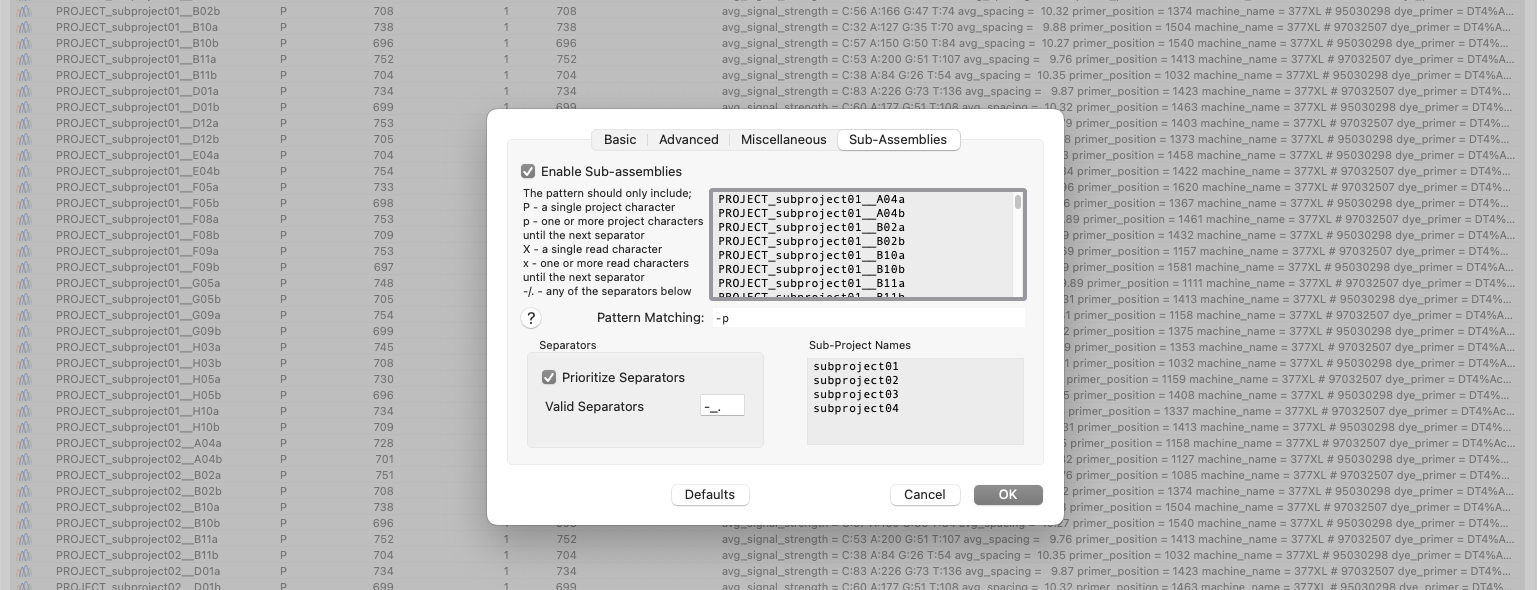
Automatic Assembly of Sub-projects with Phrap (Sub-Assemblies)
New to MacVector 18.6 is the ability to sort and assemble reads from different datasets into individual sub-projects. This functionality is located in the phrap parameters dialog. When enabled and configured appropriately for your dataset it will automatically break out the input reads into sub-projects to be assembled separately. A simple pattern-matching text box lets…
-
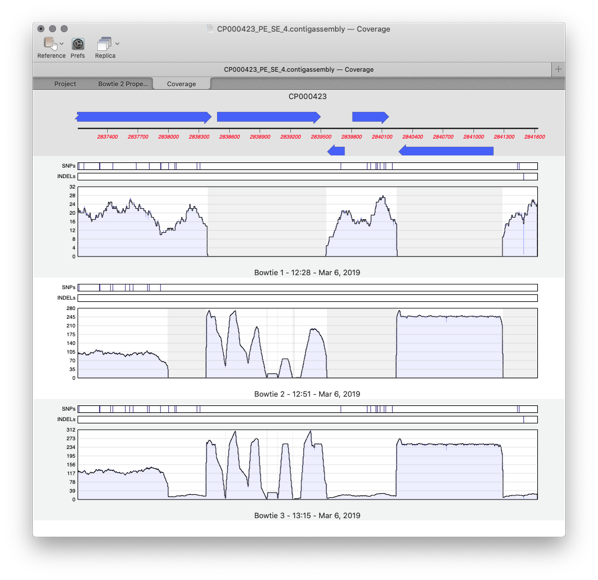
Importing Sequencher project files into MacVector
Assembler is a plugin for MacVector that provides comprehensive sequence assembly functionality. Assembler is fully integrated into MacVector and allows you to manage sequencing data with the familiar MacVector style. You can design primers directly on a contig or BLAST that contig to identify it. MacVector’s Assembly Project manager Like Sequencher, MacVector with Assembler has…
-
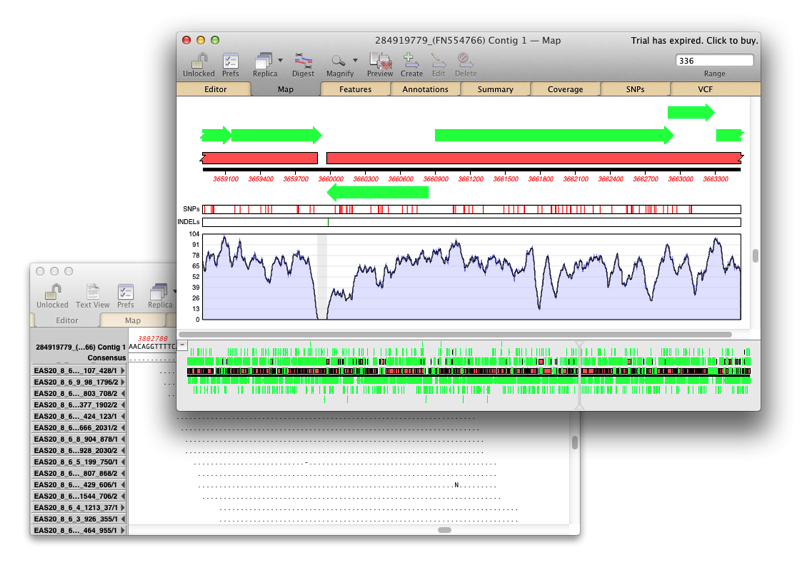
Sequence Assembly: What can Assembler do for my lab?
Assembler is fully integrated into MacVector and allows you to manage sequencing data with the familiar MacVector ease. de novo sequence assembly using Phrap, Velvet and SPAdes with Flye for PacBio and Oxford Nanopore. Reference Sequence Assembly: Map millions of reads against genomes, transcriptomes or other reference sequences using Bowtie2. Compare Genomes: Compare two related annotated…
