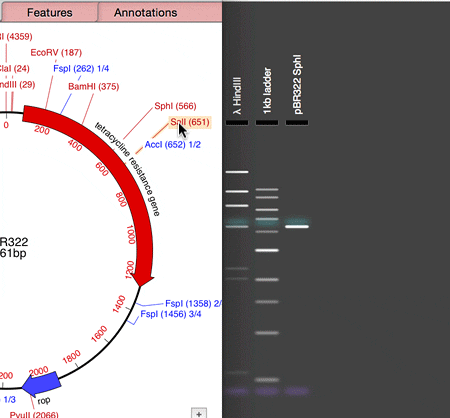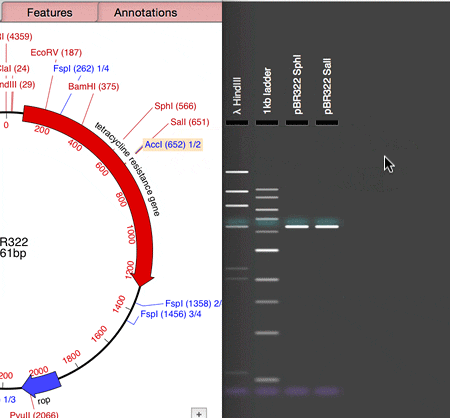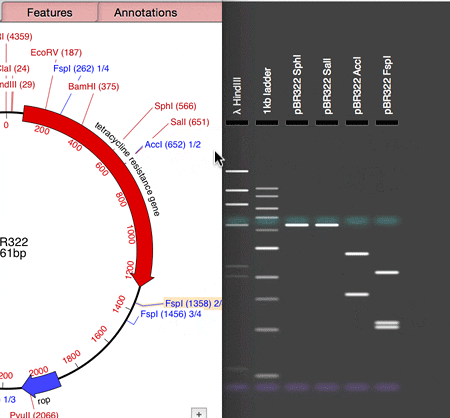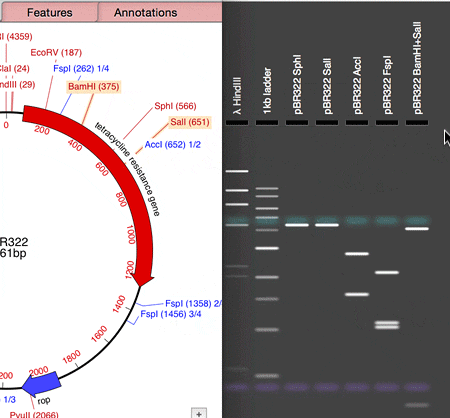
MacVector 14.5 has a Agarose Gel interface which allows you to view photo-realistic recreations of restriction digests of linear and circular DNA molecules. The gels look so realistic that users have had a hard time telling photos of their own digests from the simulation in MacVector. When you first use the new tool and compare it to your real gels, you’ll see what they mean!
The new tool makes designing digests for checking constructs very easy and quick. For example, to check the orientation of a cloned gene, drag a restriction site in that gene to the gel window to view the correct band pattern. Then for comparison, repeat the ligation in the incorrect orientation and drag the site again. You can even print out the gel to take into the darkroom as a guide for cutting out a band.
It’s incredibly easy to use the new functionality:
For a single digest
- FILE > NEW > AGAROSE GEL
- Open your sequence and switch to the MAP tab.
- Drag a restriction site from the Map tab and drop on the Gel window
For a double digest
- FILE > NEW > AGAROSE GEL
- Open your sequence and switch to the MAP tab.
- Select one restriction site, hold down SHIFT and select a second restriction site.
- Drag the sites from the Map tab and drop on the Gel window.
To change the gel marker
- Click the ADD MARKER toolbar button.
- Choose a new DNA marker.
To remove a lane
- Select the lane.
- Drag and drop the lane outside of the gel window.

Comments
2 responses to “Simulated Agarose Gels”
For the simulated agarose gel, could you add the possibility of adding restriction enzymes that do not cut into the DNA sequence. That would be useful for students that do restriction enzyme mapping in their practical course in order to see the supercoil, circular and linear DNA. Thanks.
Thanks for the comment. This is planned, and was originally due to appear in MacVector 14.5. However, the algorithm to show the proper migration of the three forms proved very difficult to show the correct migration patterns. It will appear in a future release.