MacVector’s History tab shows the edit history of your DNA sequences.
Some of MacVector’s editing tools will annotate every modification to your sequence. For example with the Cloning Clipboard, all cloning actions (such as ligating a digested fragment into a vector) create a /FRAG feature that records the source of the ligated fragment, the restriction enzymes used to digest it (and any end treatment such as Klenow) as well as a timestamp.
The History tab lists these “history” features along with a timestamp and the user who performed the edit thus allowing you to track the edit history of your sequences.
Initially only a few tools added edits to the History tab, but with each new release more will be added. For example our latest release, MacVector 18.8, has a new Auto-Annotate (via BLAST) tool that uses BLAST to find similar sequences with matching features. Every annotated feature is displayed in the History tab with the user, timestamp and the source sequence.
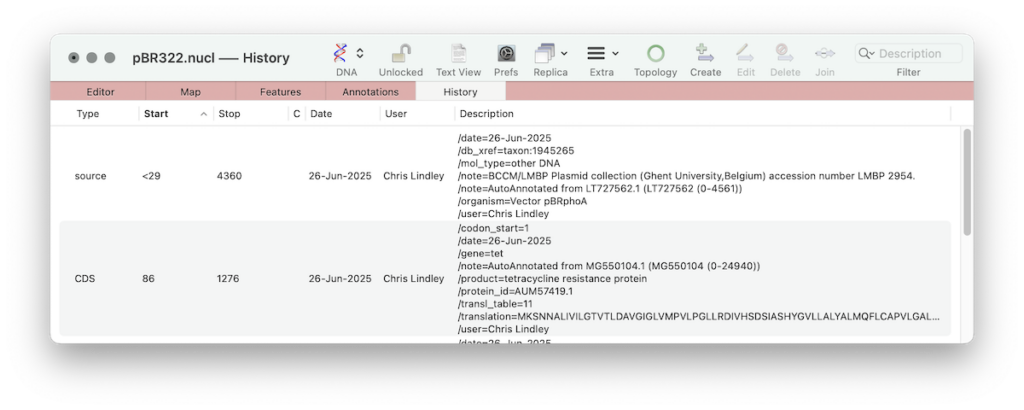
Eventually every sequence edit operation will be annotated allowing the full history of any construct to be determined and allowing a simple roll back of the construct to how it existed on a specific date. The History tab may be sorted according to the user and date as well as the usual sort columns.
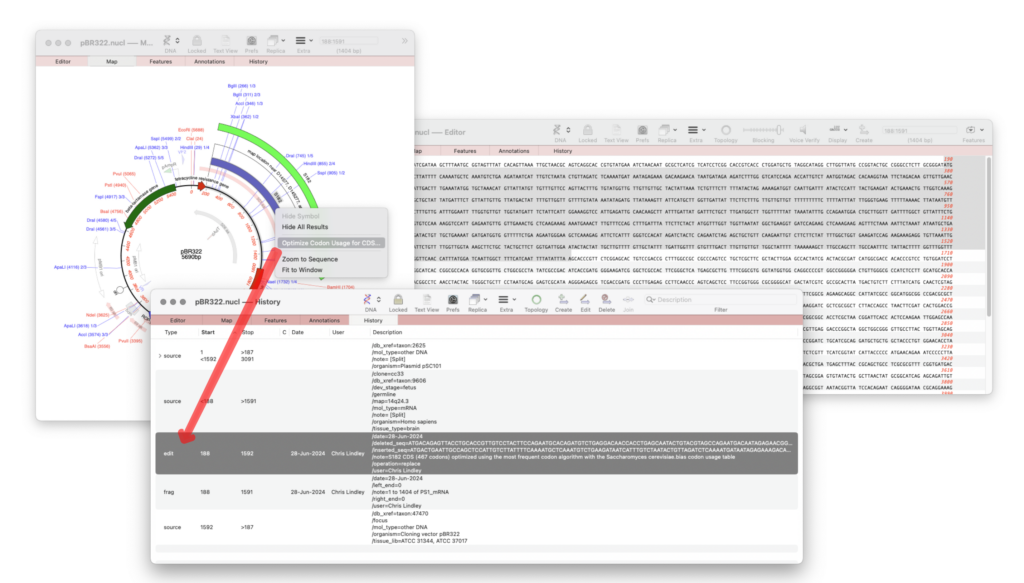
The following edit operations are shown in the History tab.
- CDS Optimization: creates an EDIT feature showing the CDS feature has been optimized.
- Cloning operations: such as ligating genes into a vector, create a FRAG feature.
- Auto-Annotate (via BLAST): Every new annotated feature has an /Action qualifier added to show how and when it was added.
- SOURCE annotations: These are a mandatory Genbank feature that summarizes the length of the sequence, scientific name of the source organism, and Taxon ID number. Can also include other information such as map location, strain, clone, tissue type, etc., if provided by submitter. Note: Source features are a standard Genbank feature and do not include timestamp or user.
Unlike all other annotation/features EDIT and FRAG features are not standard Genbank nomenclature. However, if you need to export your sequence as “pure” Genbank you can use FILE | EXPORT | Genbank Flat (strict). This will remove all non standard terms and export a file that is 100% compliant with the Genbank specification.
How to view the EDIT history of your sequence.
- Perform a ligation of a digested fragment from another sequence
- Perform a codon optimisation of a CDS feature
- Switch to the History tab of your sequence.

