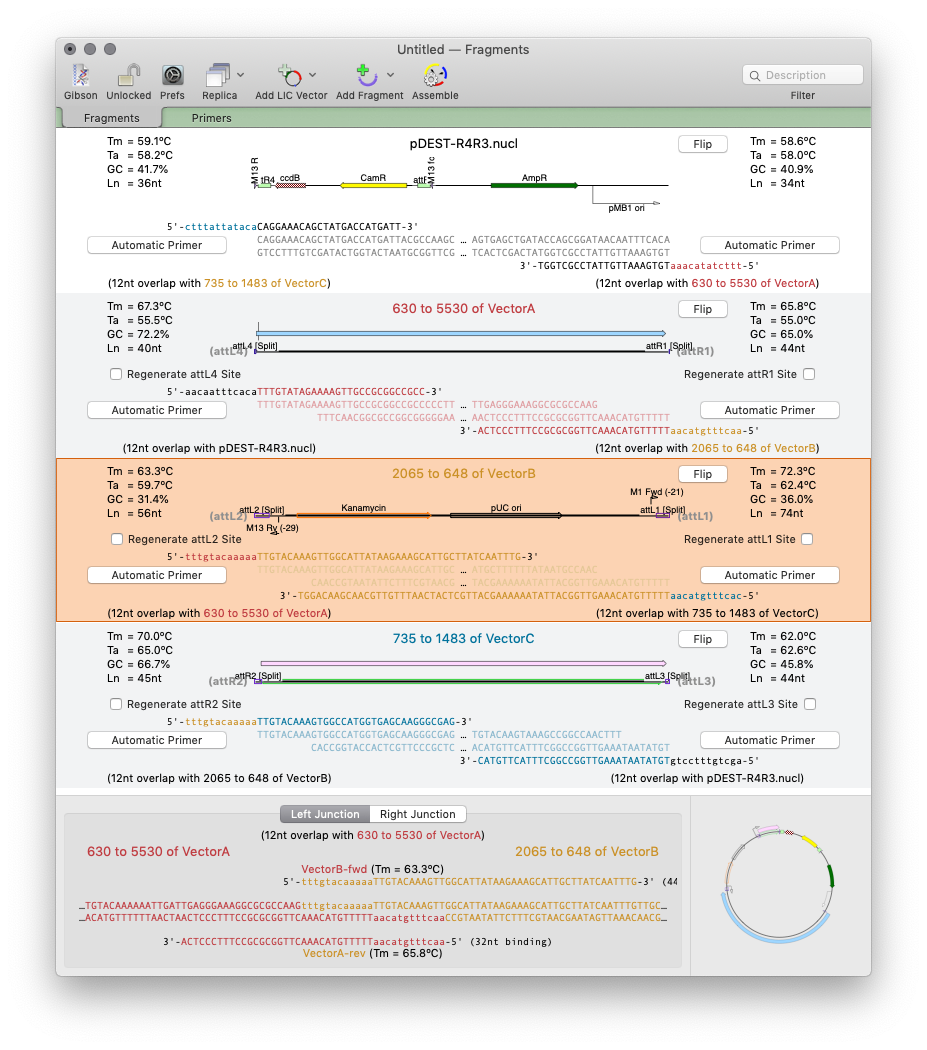
MacVector 17 has a completely new tool for automated design of ligase-independent cloning strategies. The tool supports 5’ exonuclease driven Gibson assembly as well as the T4 DNA Polymerase 3’ exonuclease “Ligase Independent Cloning” approach. MacVector can automatically design primers when you specify fragments and vectors to use. You can provide custom primers (manually or from tools such as the NEBuilder website). You can also just provide existing fragments with overlapping ends for MacVector to assemble. The interface shows the exact structure of the junctions between fragments, including relevant CDS translations for confirming the frame of protein fusions. The cloning strategy can be saved as a new cloning project document and finally automatically assembled into a single sequence for export.

MacVector 17 was released in February 2019. It contains brand new tools for comparing genomes, making restriction enzyme cloning easier, automated design of Gibson/LIC cloning strategies, highlighting assembly issues, comparing multiple sequence datasets against a single reference and making plasmid maps even more beautiful, by displaying your own primer binding sites! Finally MacVector 17 supports macOS Mojave’s Dark Mode to aid concentration on those late night primer design sessions!