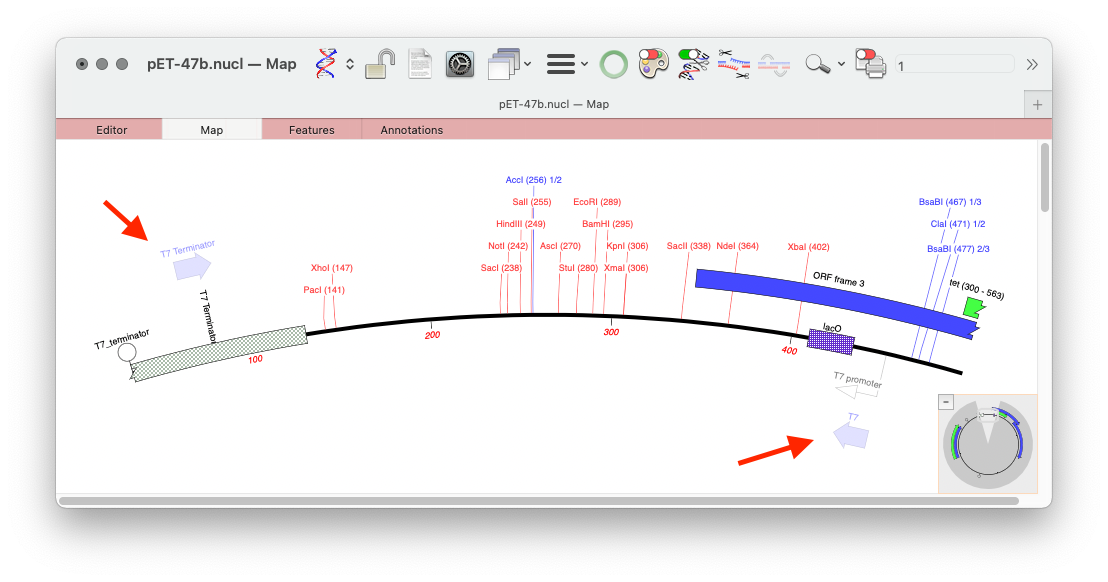
MacVector’s Scan DNA For.. tool allows you to automatically display restriction enzyme recognition sites, putative ORFs, CRISPR PAM sites, missing annotation and also it will display primer binding sites from your own Primer Database in each DNA sequence that you open. Here’s an example of a couple of primers displayed on the pET 47b LIC cloning vector on each side of the LIC cloning site. You can you quickly annotate interesting binding sites with a simple right mouse click.
The feature is controlled from the MacVector | Preferences | Scan DNA | Primers tab. My default, it uses the Primer Database.nsub file that you can find in the /Applications/MacVector/Subsequences/ folder, but you can point it to any .nsub file of your choosing.

