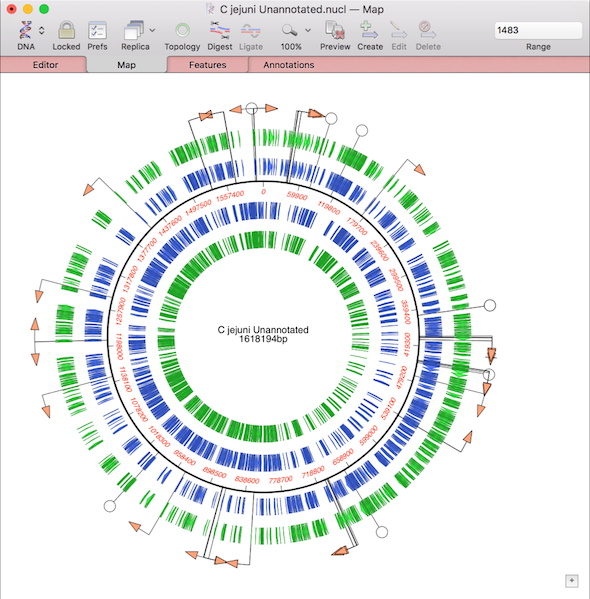
One of the more challenging aspects of dealing with large eukaryotic genomes is that most are incomplete and have many gaps. Often the genomes are available as a collection of hundreds or thousands of separate contigs, with associated annotations. This makes working with them awkward. However, increasingly, entire annotated chromosomes can be downloaded with the gaps between contigs represented by a series of Ns – typically 100. These can be downloaded from the NCBI as Genbank formatted text files which can then be imported into MacVector. However, it is not always obvious how to get entire annotated chromosomes in Genbank format. Here’s a quick primer on how to do that.
- Navigate to the Genome assembly page, e.g. Lucilia cuprina, the Australian sheep blowfly.
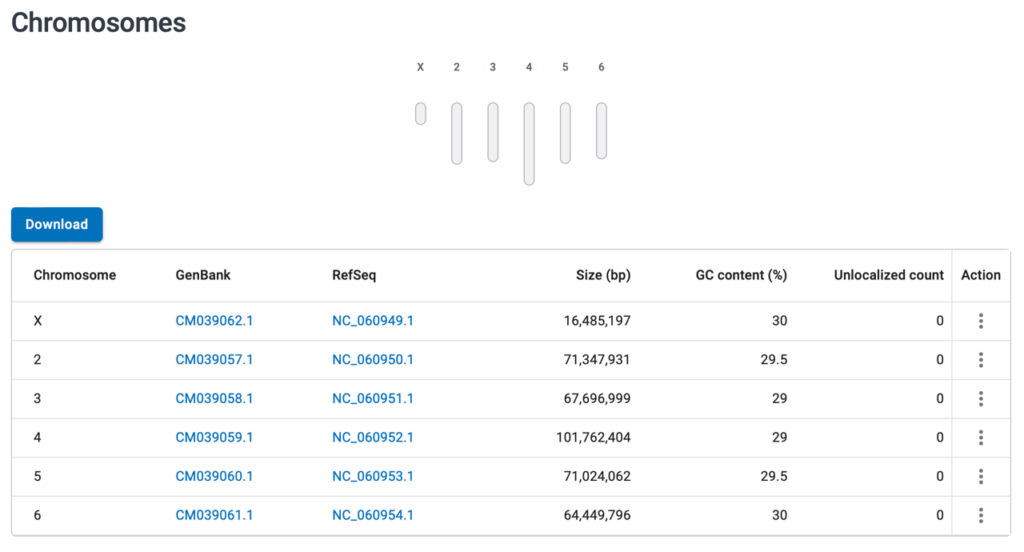
- Scroll down to the Chromosomes section. Click on either the Genbank or RefSeq link for the chromosome you are interested in. Typically, the RefSeq is fully annotated, but often the GenBank link will just point to a minimally annotated sequence.
- In the resulting window, a minimal Genbank representation of the annotations of the sequence is shown. Click on the Customize view side bar and turn on Basic Features. While you can turn on All features, often Gene, RNA and CDS are the only one’s you really care about. If the sequence is relatively short, there may also be an option to turn on the actual sequence display as well.
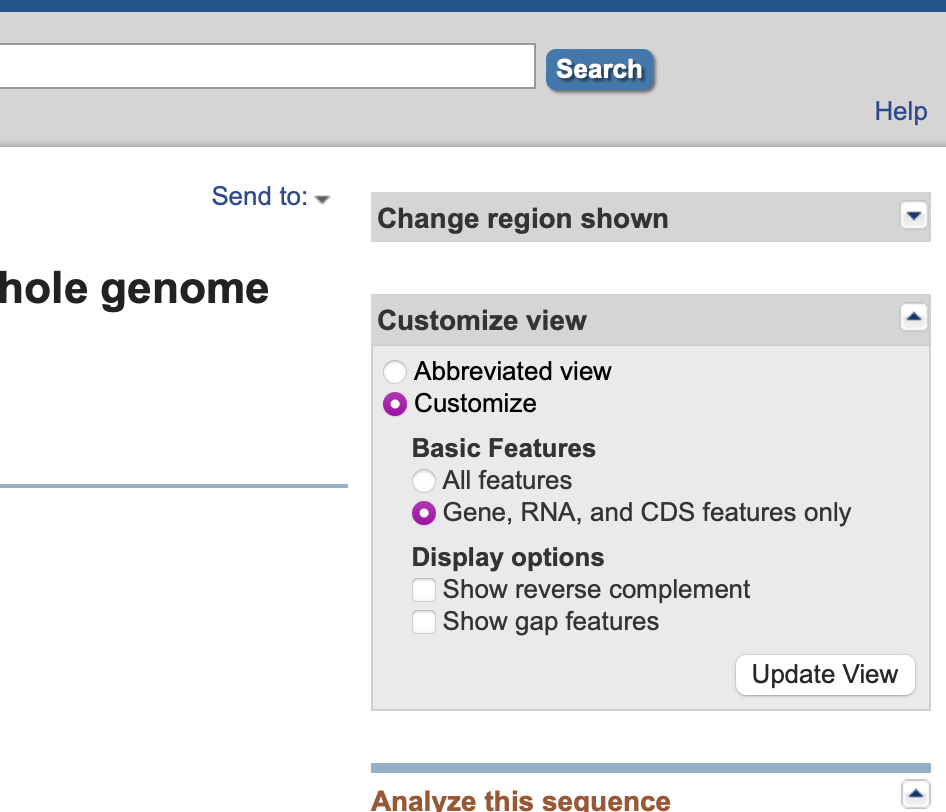
- Scroll down to ensure that you are seeing the appropriate features in the display. If you don’t see these, then the sequence is likely not annotated.
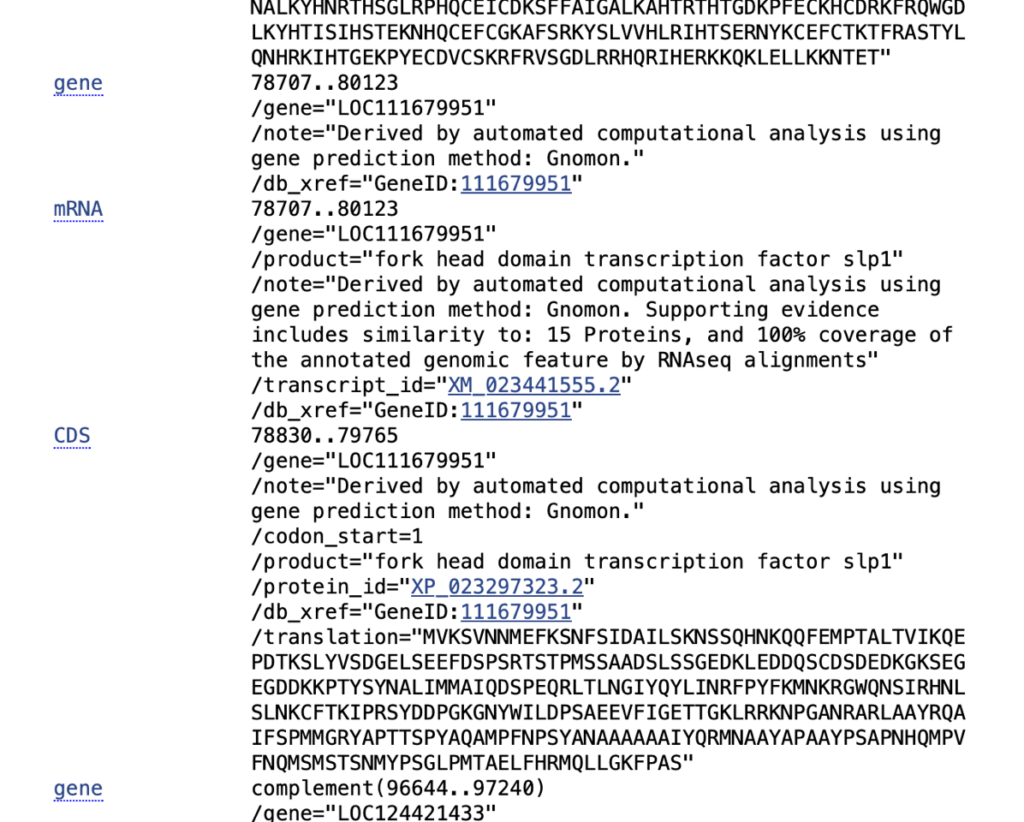
- Click on the Genbank link in the top left of the window and select GenBank (full) as the format.
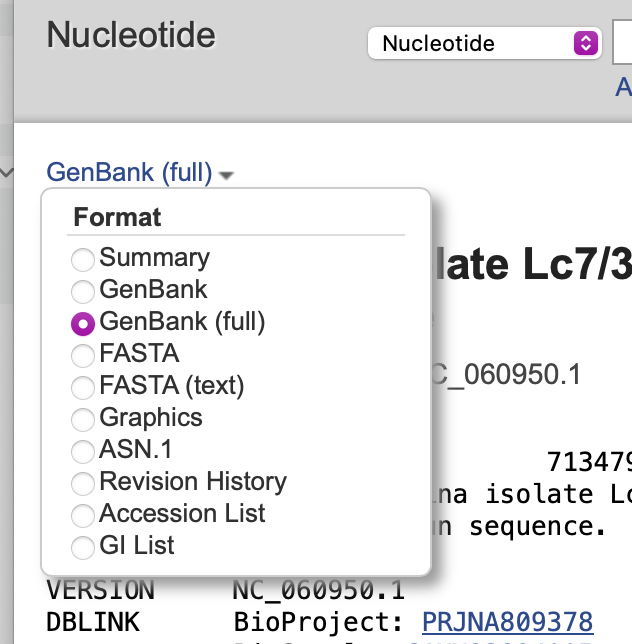
- Finally, click on the Send to: item at top right and choose Complete Record and File. The annotated sequence will be saved in your Downloads folder as “sequence.gb”. If you download additional sequences they will be named “sequence-2.gb” etc. You can now simply File | Open the file in MacVector.