New Features in MacVector 11
Gateway and TOPO Cloning Support (MacVector 11.0.3)
You can now simulate cloning manipulations using the TOPO-TA, Zero Blunt and Gateway cloning techmologies from Invitrogen. Simply open one of the pre-formatted Invitrogen vectors supplied with MacVector, (or download your own from the Invitrogen website or elsewhere) then click on the auotmatically displayed TOPO or Gateway att cloning sites to copy and paste fragments to simulate what you would do in the lab. For more detailed information, check out the tutorial.
Gateway and TOPO Cloning Tutorial
Auto Annotation.
How often have you received or downloaded a vector or other DNA sequence that has no annotations? The new auto annotation function in MacVector 11 can scan a folder full of existing annotated sequences and automatically add matching features to the bare sequence. Not only does the algorithm add the features, but it also copies the appearance information of the matching features so you can be assured that e.g. an ampicillin resistance gene is always a blue arrow, an M13 origin is always a striped red box etc.
Click Cloning Enhancements
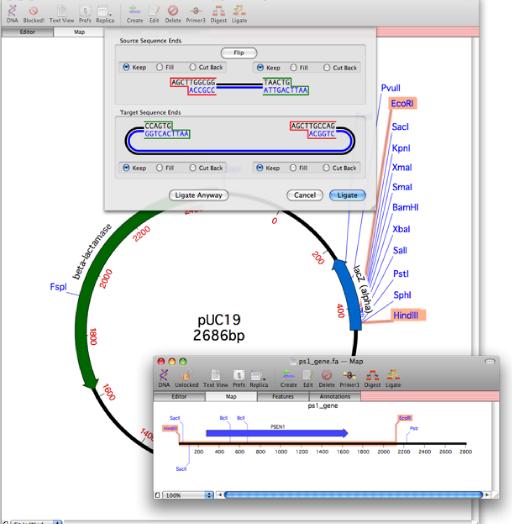
In MacVector 11 you can manipulate the ends of restriction fragments before joining them together. A new interface lets you cut back or fill in each end of the source or target molecule before ligation.
Floating Analysis Toolbar.
For those users who prefer to click on toolbar buttons to initiate analyses, MacVector 11 introduces a new floating toolbar window containing buttons for all common MacVector analysis functions. You can customize the toolbar to show just the functions you use most often or show them all for rapid access to every available algorithm. You can also add analysis buttons to the toolbars of normal sequence windows for ready access to commonly used functions.
Sequence Editor Changes
The primary sequence editor has been rewritten using modern OS X code to better handle long sequences and to provide an OS X look and feel. In addition, you can now display 3 or 6 frame translations below the sequence that update instantly as you edit the sequence.
Next Generation Sequencing
The optional add-on assembler module has been enhanced to provide support for next generation sequencing machines. Short read data may be imported in Fastq format and assembled them using the latest version of phrap.
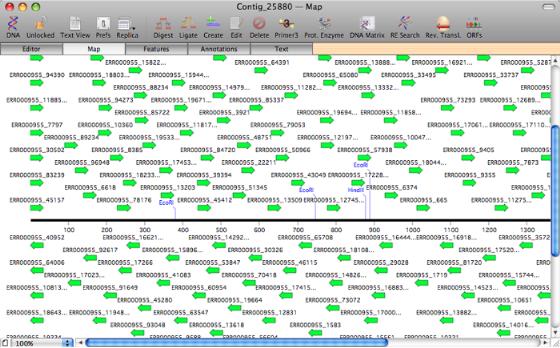
A Map view of a contig assembled from Solexa reads imported from a Fastq file.
Miscellaneous Enhancements.
As always, we add a slew of minor enhancements to each release of MacVector designed to improve workflows, speed up processing or provide better integration with the operating system. Look for the ability to change the colors of chromatogram file displays for red/green color blind users, improved import of sequences from Vector NTI and better handling of genomic sequences.
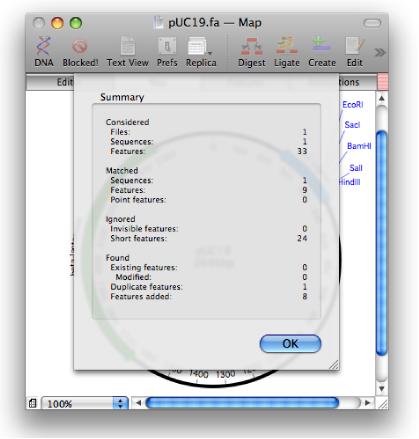
MacVector 11 will let you annotate blank sequences using a library of curated annotated sequences.
Availability
MacVector 11 is available now!. Click here to sign up to download a fully functional 21-day trial.
|

2x.png)


2x.png)
