What's New in MacVector 15?
Overview
There are two main new features in MacVector 15, both targeted at users with an interest in proteins. First, MacVector can now connect to the online InterProScan database to help you identify functional domains in your protein. Second, the multiple sequence alignment interface has been completely rewritten to let you align DNA sequences via their amino acid translations and also to let you define a reference sequence when aligning proteins. As always, there are a multitude of minor enhancements and bug fixes designed to improve your everyday experience using MacVector. You can view the release notes here.
InterProScan
You can now scan any protein for functional domains using the Database | Internet InterProScan Search... menu item. When the search completes, you get a graphical result window like this;
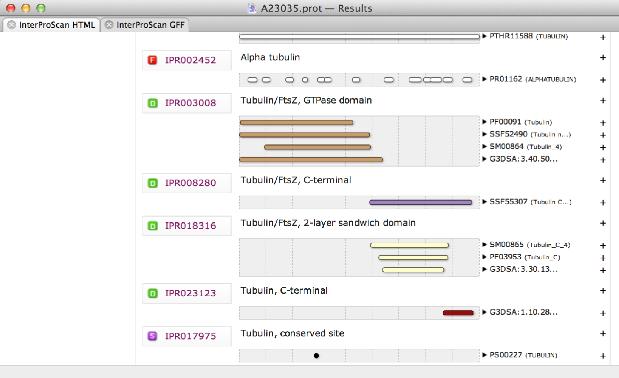
You can make any match a permanent annotation of your starting protein sequence by simply clicking on the "+" icon next to the entry.
MSA Enhancements
The Multiple Sequence Alignment window has had a number of enhancements. Most are accessed through a new "Mode" button on the Editor tab toolbar.
Aligning DNA via Translations - if you open or create a Nucleic Acid alignment, there are new display modes;
NA - this mode is exactly the same as earlier versions - all of the display, editing and alignment tools use the nucleic acid sequences, just as you would expect.
virtualAA - this mode translates the NA sequences (using the currently selected Genetic Code) and displays the translations just as if this was a protein alignment. DNA is always translated starting at the first residue. If you run an alignment algorithm while this mode is selected, the alignment will use the amino acid sequences to align the sequences. You can always toggle the mode back to NA to view the nucleic acid sequences after the alignments. In this mode you have minimal editing capabilities - you can "slide" amino acids and delete them, but most functions are disabled because there are NA sequences that underlie the amino acids, so changing/adding amino acids would lead to ambiguous nucleic acid residues.
NA+virtualAA - this mode displays both the nucleic acid and amino acid sequences. Here you can edit the nucleic acid residues just as you would in NA mode and immediately see the effect of any edits on the translations. However, any ClustalW/Muscle/T-Coffee alignments will still use the translated amino acid sequences to align the sequences, so you get the best of both approaches;
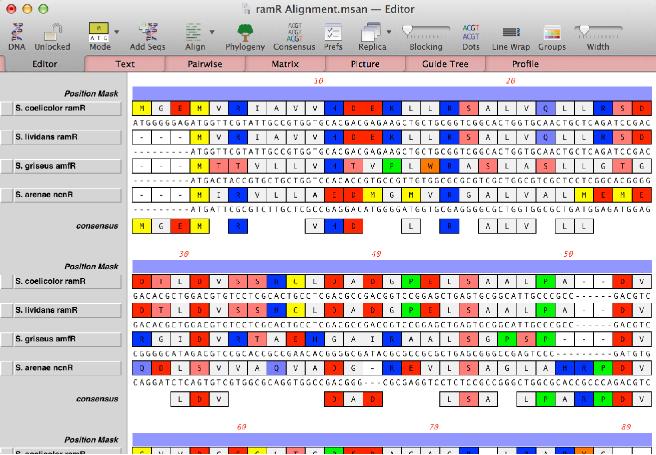
Designating a reference -Normally, the multiple sequence alignment interface treats all of the sequences equally and displays the consensus below the alignment. MacVector 15 allows you to designate one sequence as the reference. This is also accessed through the new Mode toolbar button. Practically, this is most useful for protein sequences. If you want to align DNA sequences to a reference you should be using the Analyze | Align To Reference functionality. However, for protein sequences, this is a great way to view the alignments in such a way that you can easily visualize the differences between a set of proteins and a reference protein. To invoke this mode, simply drag your preferred reference protein to the bottom of the alignment and then select Show Reference and the last sequence will be turned into a reference and will replace the consensus. Here we see an alignment where matches to the reference are shown by dots;
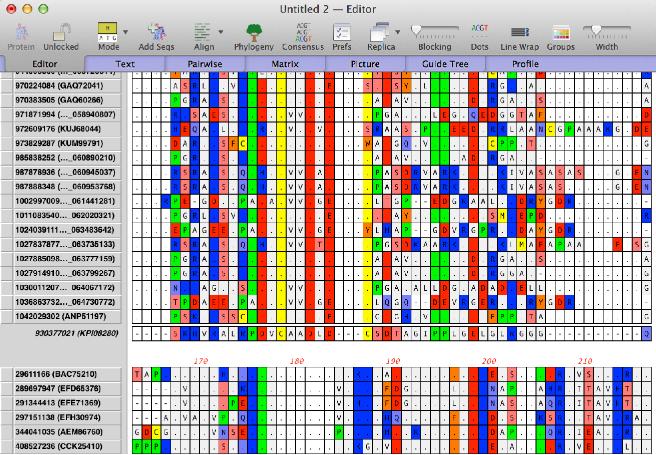
|

2x.png)


2x.png)
