There are many output windows throughout MacVector that display aligned sequences. If you run an Align To Folder, Create Dotplot or Internet Blast Search, one or more of the output windows will show alignments in a plain text format with some sort of character indicating matching residues. The default is to use the vertical | character for matching residues;
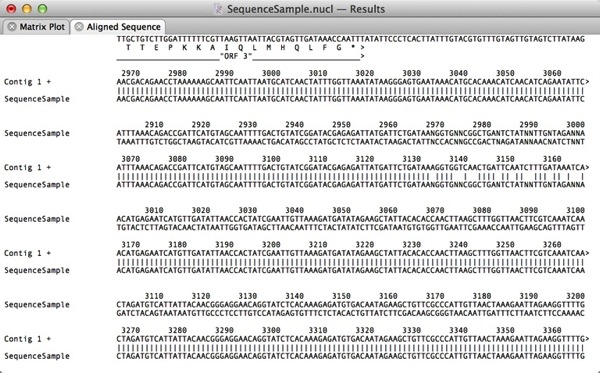
You can change the characters used in these displays using the MacVector | Preferences | Aligned Sequence pane. Note that these fields just accept a single character to be used in the alignment display;
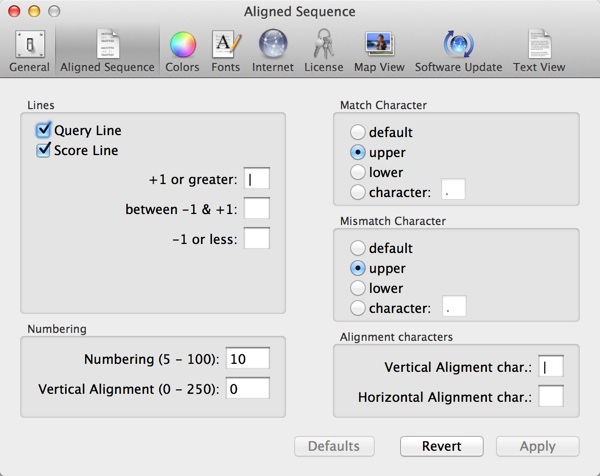
The “+1 or greater” field indicates the character used for matching characters using the currently selected scoring matrix. For DNA, this usually indicates identical matches, but for protein sequences this can be certain similar residues (leucine versus isoleucine for example).
The “between -1 & +1” field is used for “neutral” matches – e.g. an “N” against a “G” for DNA. By default this is set to a space character.
Finally, the “-1 or less” character is used for mismatches.
Sometimes, you may be comparing sequences where you are far more interested in the mismatches between them than the matches. In the example above, there are mismatches in one section that do not immediately leap out form the page. Lets see what happens if we change the characters from the default to this;
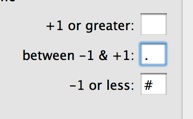
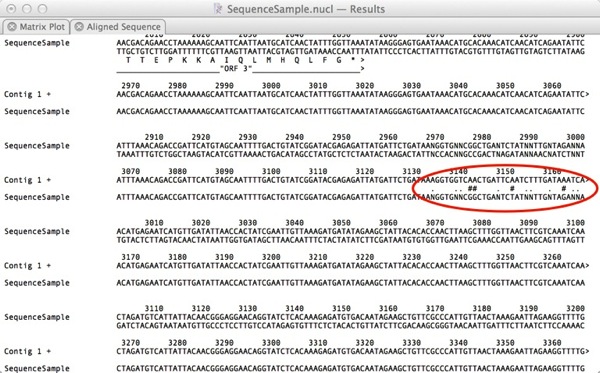
After we click Apply, the alignment text window updates to make it much clearer where the mismatches lie;
This is an article in a long running series of tips to help you get the most out of MacVector. If you want to get notified every time a new tip gets published, follow us @MacVector on twitter (or check the feed for the hashtag #101MacVectorTips) or like us on Facebook.