The NCBI Sequence Read Archive (SRA) database is a huge resource of Next Generation Sequencing experimental data. Many groups and laboratories deposit data here that they have generated for their own specific projects that can be datamined for other unrelated projects with a minimum of effort.
MacVector contains a number of powerful tools that can be used to extract and analyze specific sequences from large quantities of NGS data. We recently used these tools to clone the sequence of 19 distinct C2H2 Zinc Finger proteins from NGS RNA-Seq data prepared from root tissue of the Aloe vera plant.
The basic steps to do this were;
- Use Align to Folder to find and extract all pairs of reads that could potentially encode the conserved QALGGH domain from C2H2 Zn Finger proteins
- Assemble the reads using phrap, velvet and/or SPAdes to generate multiple contigs
- Analyze contigs to identify and translate protein-coding ORFs
- Extend contigs when required using additional rounds of Align to Folder, contig assembly and Align to Reference
- Annotate proteins using the built-in InterProScan function
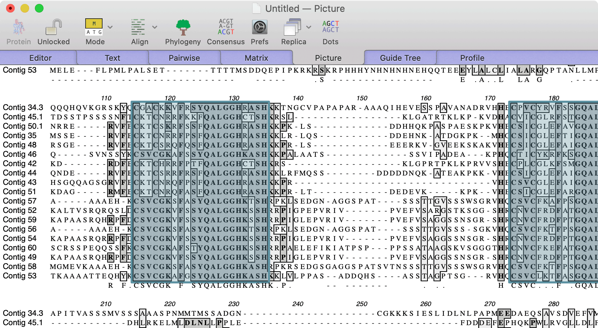
- Align proteins using ClustalW and visualize the shared QALGGH domains
The full tutorial is available as a PDF and the required data files are also available to download direct from the SRA.