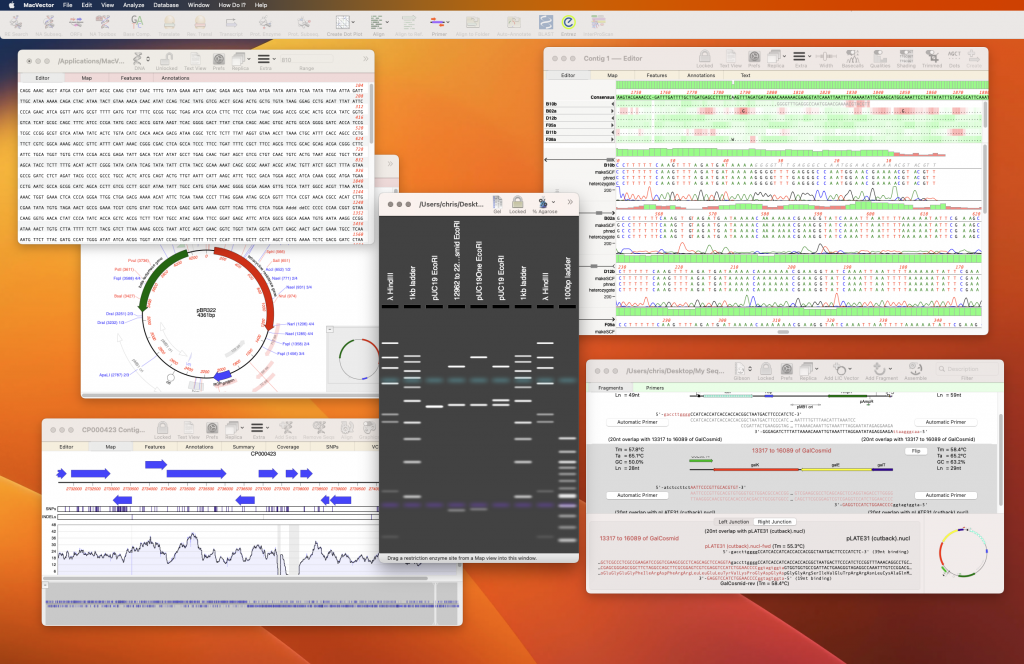
MacVector 18.6 is the latest release of the best sequence analysis application for the Mac. MacVector 18.6 is a Universal Binary application and will run natively on both Apple Silicon Macs and Intel Macs.
We think there is no better tool for designing primers, preparing beautiful plasmid maps, comparing sequences, CRISPR INDEL analysis, Gibson Assembly, LIC and restriction enzyme cloning workflows, comparing genomes, sequence assembly and much more on the Mac.
MacVector 18.6 is fully supported on macOS High Sierra to macOS Ventura.
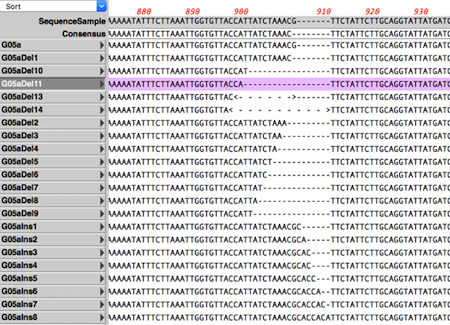
CRISPR Indel Analysis: following CRISPR editing of a target. Compare Genomes: identify identical, similar and weakly similar genes. de novo sequence assembly using Phrap, Velvet and SPAdes with Flye for PacBio and Oxford Nanopore. Reference sequence assembly using Bowtie. Sequencher compatible: Directly import Sequencher assemblies into MacVector’s Assembly Projects. Automatic Assembly of Sub-projects with Phrap: Automatically break out reads into sub-projects to be assembled separately. RNA-Seq analysis Read depth visualization with per gene coverage data (RPKM & TPM). Circularize genomes: toolkit for finishing assemblies and circularizing bacterial ones. 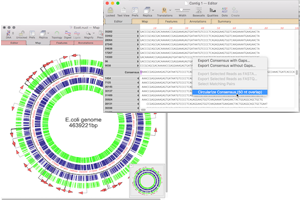
Sequence confirmation: quickly verify sequenced constructs. Heterozygote analysis: report on possible heterozygotes and change the sequence with an IUPAC ambiguity code. Coverage Tab: compare multiple assemblies with expression level comparison. Outline shared aligned domains: Multiple sequence alignments display shared domains. 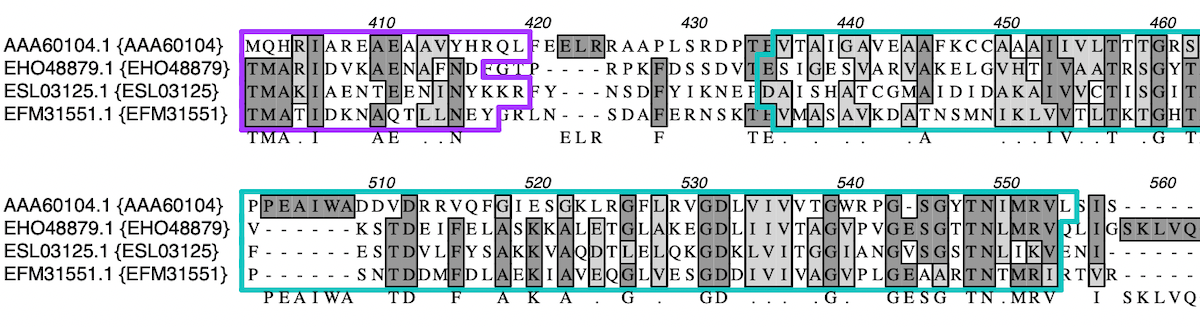
Easily create beautiful plasmid maps Automatically annotate missing features, primer binding sites CRISPR PAM sites and putative ORFs. InterProScan: Scan proteins for functional domains. |
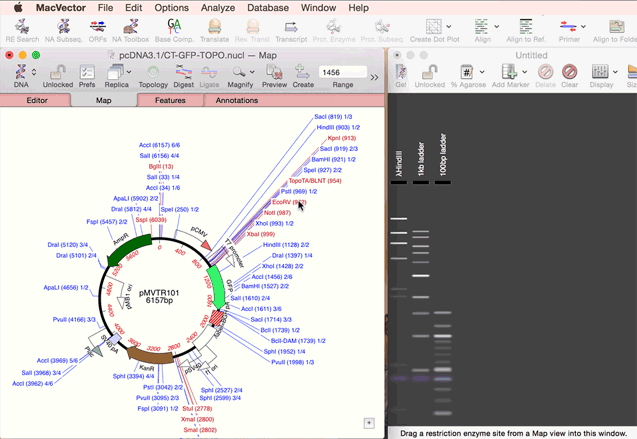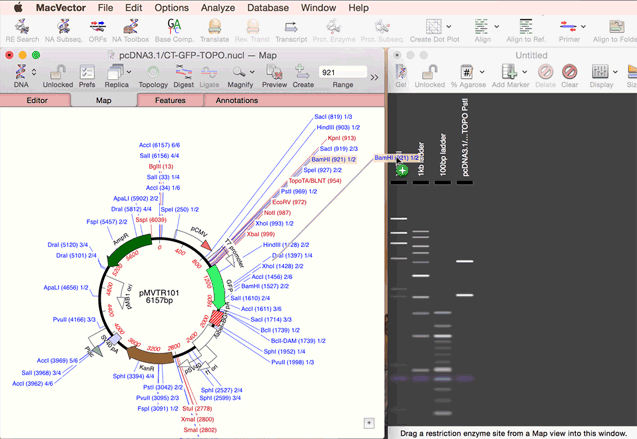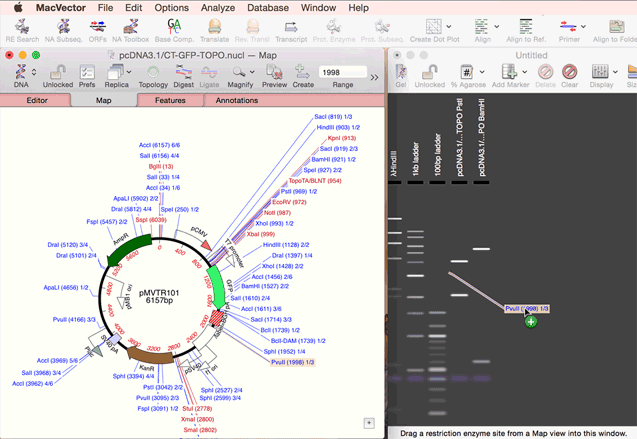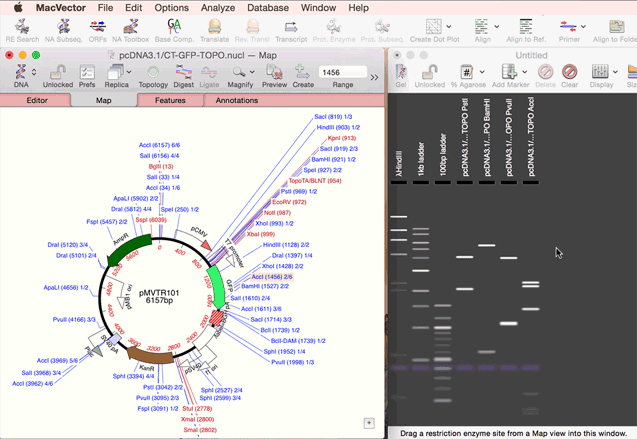
As simple as dragging a fragment to a cloning vector. Cloning history Every digestion and ligation is documented. Cloning Clipboard Store your digested fragments for cloning into a vector. 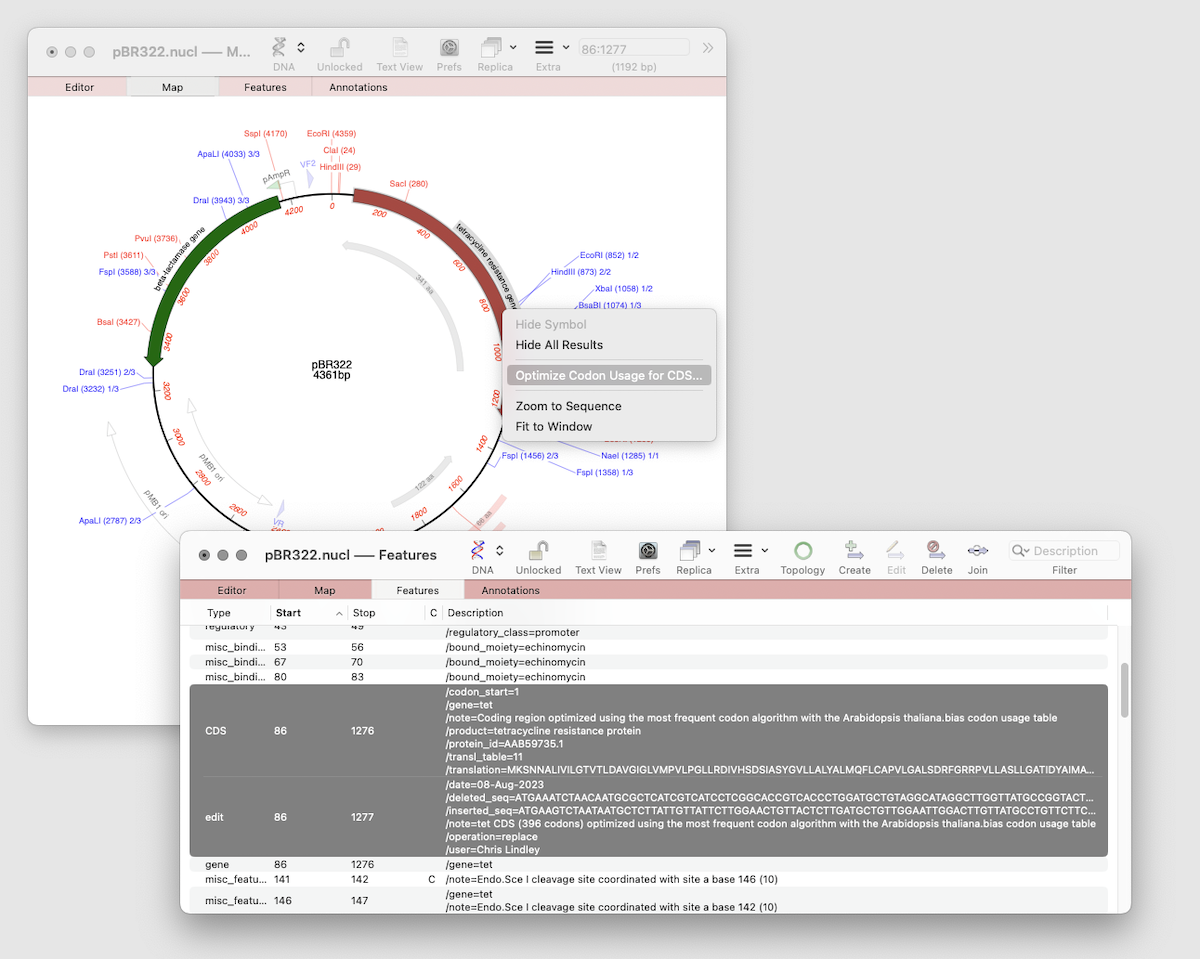
One step codon usage optimization: for enhanced expression of a CDS in a different organism. Simulated Agarose Gel: identify restriction sites to screen minipreps. Restriction Enzyme Picker: easy identification of useful enzyme cut sites. 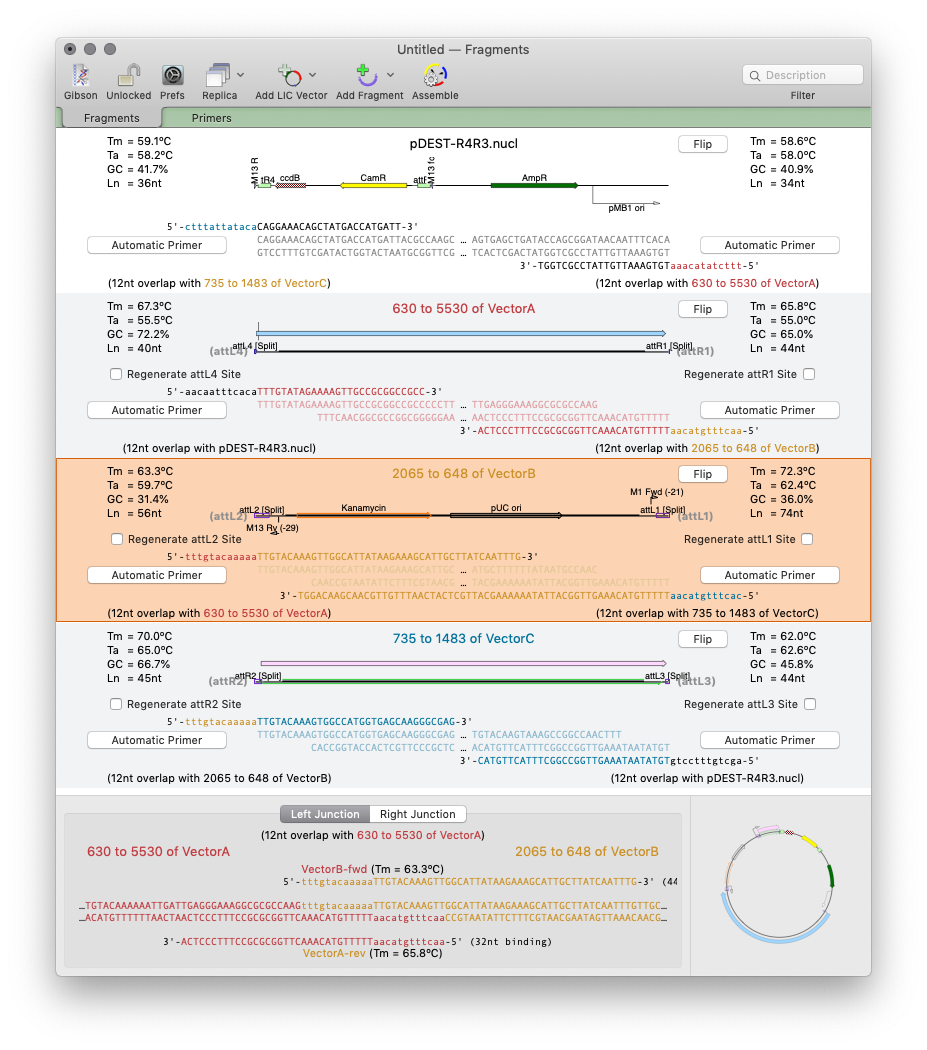
Gibson/Ligase-Independent Assemblies: project based design of workflows inc. primer design. QuickTest Primer makes primer design easy! Slide your primer along the template until it aligns perfectly! Add tails to your primers with silent restriction sites/mismatches and view reading frame changes. Quickly design pairs of primers click a region to get the best primer pairs to amplify it. Scan For… Missing Primers: automatically display binding sites from your lab’s primer collection |