When analyzing/assembling/aligning NGS data, there are many scenarios where you might want to separate out the reads representing different genotypes or variant sequences. MacVector makes this very easy. Take a reference sequence and choose Analyze->Align to Reference. Now click the Add Seqs button and select and add your NGS data files. NOTE: if your reference represents just a subset of the data in the NGS files, you might want to first filter the data using Align to Folder
Here we see an Align to Reference where about half the reads have obvious SNPs compared to the reference. Note that the Dots toolbar button is toggled on to help emphasize the mismatches;
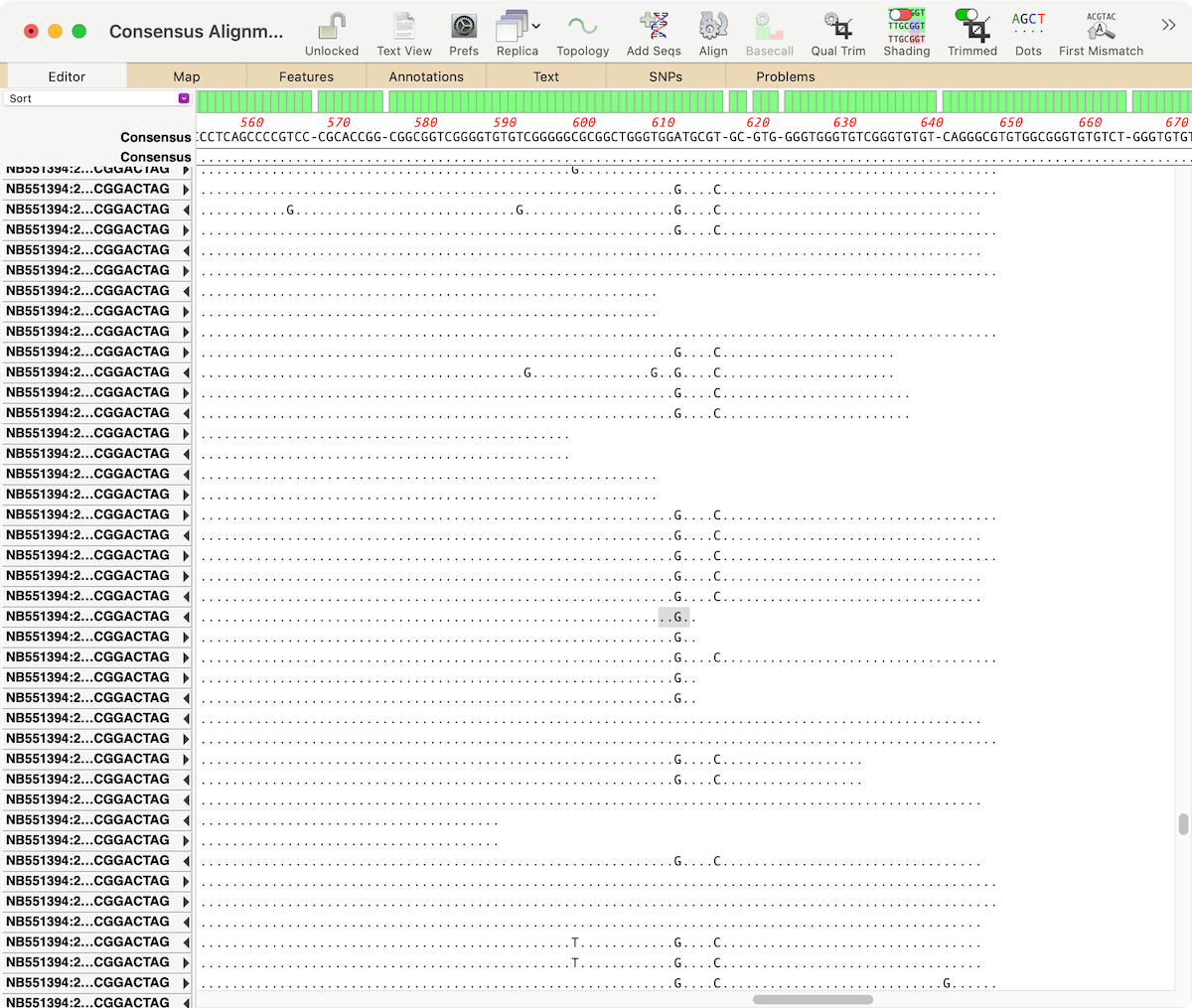
To select all of the reads that contain the SNP, first select a few residues around that SNP, as shown above. This helps ignore the occasional “bad” sequence, though, for most purposes, you can just select the one residue. Then right-click ([ctrl]
If your sequence has multiple SNPs/genotypes/repeats, you can always then choose the right-click Delete Selected Reads option to remove those reads and start again on another set.